A Cancer Researcher Opens Up About His Astonishing Breakthrough
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.
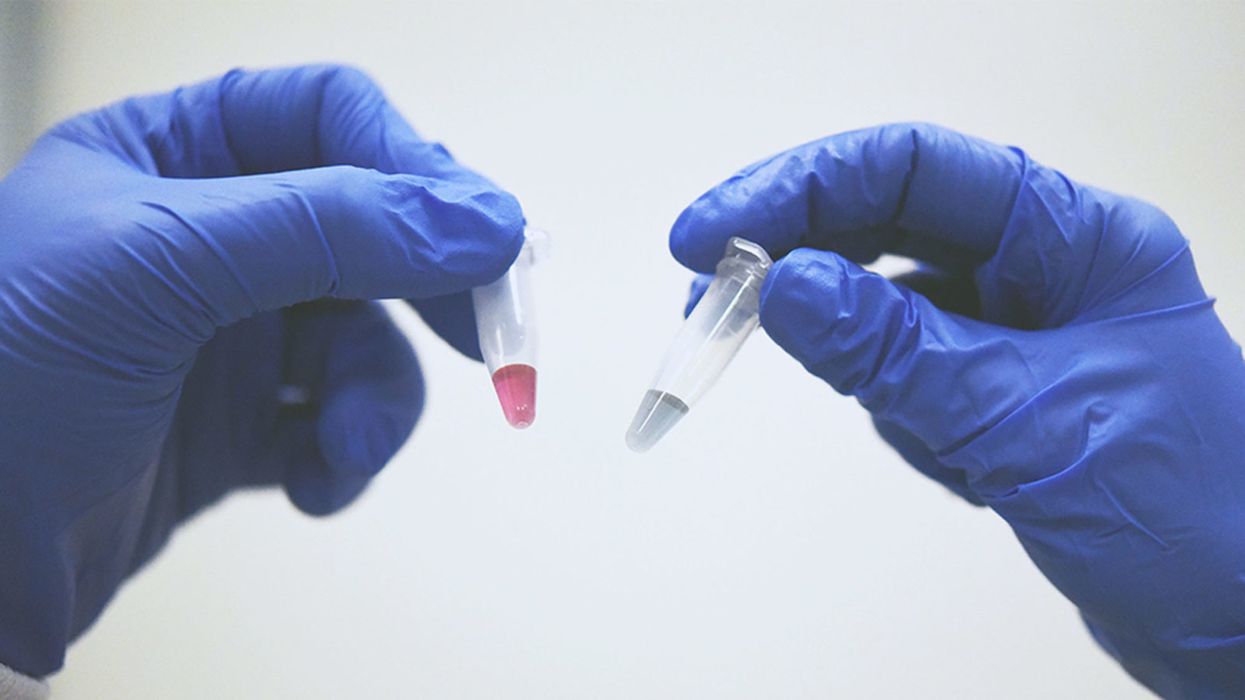
A simple ten-minute universal cancer test that can be detected by the human eye or an electronic device - published in Nature Communications (Dec 2018) by the Trau lab at the University of Queensland. Red indicates the presence of cancerous cells and blue doesn't.
Matt Trau, a professor of chemistry at the University of Queensland, stunned the science world back in December when the prestigious journal Nature Communications published his lab's discovery about a unique property of cancer DNA that could lead to a simple, cheap, and accurate test to detect any type of cancer in under 10 minutes.
No one believed it. I didn't believe it. I thought, "Gosh, okay, maybe it's a fluke."
Trau granted very few interviews in the wake of the news, but he recently opened up to leapsmag about the significance of this promising early research. Here is his story in his own words, as told to Editor-in-Chief Kira Peikoff.
There's been an incredible explosion of knowledge over the past 20 years, particularly since the genome was sequenced. The area of diagnostics has a tremendous amount of promise and has caught our lab's interest. If you catch cancer early, you can improve survival rates to as high as 98 percent, sometimes even now surpassing that.
My lab is interested in devices to improve the trajectory of cancer patients. So, once people get diagnosed, can we get really sophisticated information about the molecular origins of the disease, and can we measure it in real time? And then can we match that with the best treatment and monitor it in real time, too?
I think those approaches, also coupled with immunotherapy, where one dreams of monitoring the immune system simultaneously with the disease progress, will be the future.
But currently, the methodologies for cancer are still pretty old. So, for example, let's talk about biopsies in general. Liquid biopsy just means using a blood test or a urine test, rather than extracting out a piece of solid tissue. Now consider breast cancer. Still, the cutting-edge screening method is mammography or the physical interrogation for lumps. This has had a big impact in terms of early detection and awareness, but it's still primitive compared to interrogating, forensically, blood samples to look at traces of DNA.
Large machines like CAT scans, PET scans, MRIs, are very expensive and very subjective in terms of the operator. They don't look at the root causes of the cancer. Cancer is caused by changes in DNA. These can be changes in the hard drive of the DNA (the genomic changes) or changes in the apps that the DNA are running (the epigenetics and the transcriptomics).
We don't look at that now, even though we have, emerging, all of these technologies to do it, and those technologies are getting so much cheaper. I saw some statistics at a conference just a few months ago that, in the United States, less than 1 percent of cancer patients have their DNA interrogated. That's the current state-of-the-art in the modern medical system.

Professor Matt Trau, a cancer researcher at the University of Queensland in Australia.
(Courtesy)
Blood, as the highway of the body, is carrying all of this information. Cancer cells, if they are present in the body, are constantly getting turned over. When they die, they release their contents into the blood. Many of these cells end up in the urine and saliva. Having technologies that can forensically scan the highways looking for evidence of cancer is little bit like looking for explosives at the airport. That's very valuable as a security tool.
The trouble is that there are thousands of different types of cancer. Going back to breast cancer, there's at least a dozen different types, probably more, and each of them change the DNA (the hard drive of the disease) and the epigenetics (or the RAM memory). So one of the problems for diagnostics in cancer is to find something that is a signature of all cancers. That's been a really, really, really difficult problem.
Ours was a completely serendipitous discovery. What we found in the lab was this one marker that just kept coming up in all of the types of breast cancers we were studying.
No one believed it. I didn't believe it. I thought, "Gosh, okay, maybe it's a fluke, maybe it works just for breast cancer." So we went on to test it in prostate cancer, which is also many different types of diseases, and it seemed to be working in all of those. We then tested it further in lymphoma. Again, many different types of lymphoma. It worked across all of those. We tested it in gastrointestinal cancer. Again, many different types, and still, it worked, but we were skeptical.
Then we looked at cell lines, which are cells that have come from previous cancer patients, that we grow in the lab, but are used as model experimental systems. We have many of those cell lines, both ones that are cancerous, and ones that are healthy. It was quite remarkable that the marker worked in all of the cancer cell lines and didn't work in the healthy cell lines.
What could possibly be going on?
Well, imagine DNA as a piece of string, that's your hard drive. Epigenetics is like the beads that you put on that string. Those beads you can take on and off as you wish and they control which apps are run, meaning which genetic programs the cell runs. We hypothesized that for cancer, those beads cluster together, rather than being randomly distributed across the string.
Ultimately, I see this as something that would be like a pregnancy test you could take at your doctor's office.
The implications of this are profound. It means that DNA from cancer folds in water into three-dimensional structures that are very different from healthy cells' DNA. It's quite literally the needle in a haystack. Because when you do a liquid biopsy for early detection of cancer, most of the DNA from blood contains a vast abundance of healthy DNA. And that's not of interest. What's of interest is to find the cancerous DNA. That's there only in trace.
Once we figured out what was going on, we could easily set up a system to detect the trace cancerous DNA. It binds to gold nanoparticles in water and changes color. The test takes 10 minutes, and you can detect it by eye. Red indicates cancer and blue doesn't.
We'revery, very excited about where we go from here. We're starting to test the test on a greater number of cancers, in thousands of patient samples. We're looking to the scientific community to engage with us, and we're getting a really good response from groups around the world who are supplying more samples to us so we can test this more broadly.
We also are very interested in testing how early can we go with this test. Can we detect cancer through a simple blood test even before there are any symptoms whatsoever? If so, we might be able to convert a cancer diagnosis to something almost as good as a vaccine.
Of course, we have to watch what are called false positives. We don't want to be detecting people as positives when they don't have cancer, and so the technology needs to improve there. We see this version as the iPhone 1. We're interested in the iPhone 2, 3, 4, getting better and better.
Ultimately, I see this as something that would be like a pregnancy test you could take at your doctor's office. If it came back positive, your doctor could say, "Look, there's some news here, but actually, it's not bad news, it's good news. We've caught this so early that we will be able to manage this, and this won't be a problem for you."
If this were to be in routine use in the medical system, countless lives could be saved. Cancer is now becoming one of the biggest killers in the world. We're talking millions upon millions upon millions of people who are affected. This really motivates our work. We might make a difference there.
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.
A robot server, controlled remotely by a disabled worker, delivers drinks to patrons at the DAWN cafe in Tokyo.
A sleek, four-foot tall white robot glides across a cafe storefront in Tokyo’s Nihonbashi district, holding a two-tiered serving tray full of tea sandwiches and pastries. The cafe’s patrons smile and say thanks as they take the tray—but it’s not the robot they’re thanking. Instead, the patrons are talking to the person controlling the robot—a restaurant employee who operates the avatar from the comfort of their home.
It’s a typical scene at DAWN, short for Diverse Avatar Working Network—a cafe that launched in Tokyo six years ago as an experimental pop-up and quickly became an overnight success. Today, the cafe is a permanent fixture in Nihonbashi, staffing roughly 60 remote workers who control the robots remotely and communicate to customers via a built-in microphone.
More than just a creative idea, however, DAWN is being hailed as a life-changing opportunity. The workers who control the robots remotely (known as “pilots”) all have disabilities that limit their ability to move around freely and travel outside their homes. Worldwide, an estimated 16 percent of the global population lives with a significant disability—and according to the World Health Organization, these disabilities give rise to other problems, such as exclusion from education, unemployment, and poverty.
These are all problems that Kentaro Yoshifuji, founder and CEO of Ory Laboratory, which supplies the robot servers at DAWN, is looking to correct. Yoshifuji, who was bedridden for several years in high school due to an undisclosed health problem, launched the company to help enable people who are house-bound or bedridden to more fully participate in society, as well as end the loneliness, isolation, and feelings of worthlessness that can sometimes go hand-in-hand with being disabled.
“It’s heartbreaking to think that [people with disabilities] feel they are a burden to society, or that they fear their families suffer by caring for them,” said Yoshifuji in an interview in 2020. “We are dedicating ourselves to providing workable, technology-based solutions. That is our purpose.”
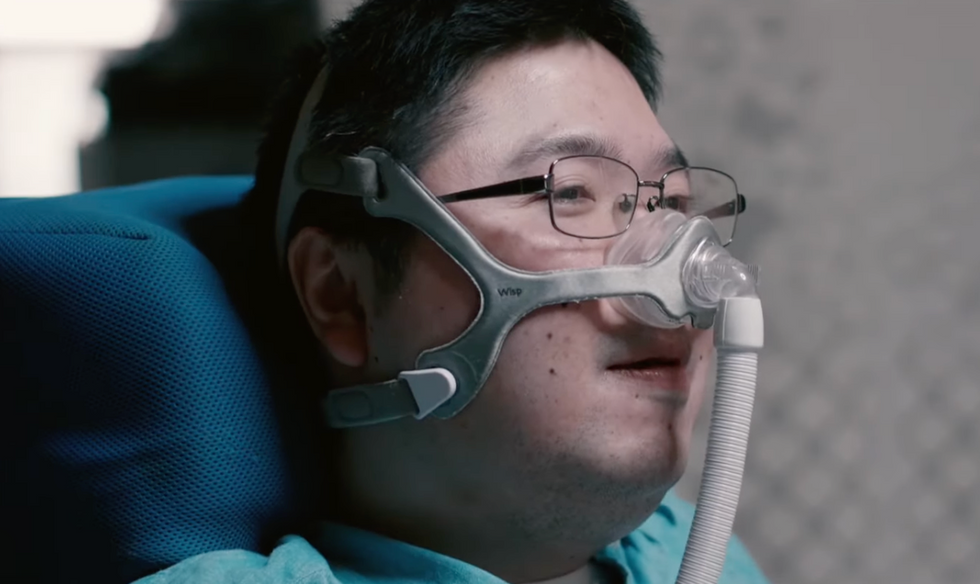
Shota, Kuwahara, a DAWN employee with muscular dystrophy, agrees. "There are many difficulties in my daily life, but I believe my life has a purpose and is not being wasted," he says. "Being useful, able to help other people, even feeling needed by others, is so motivational."
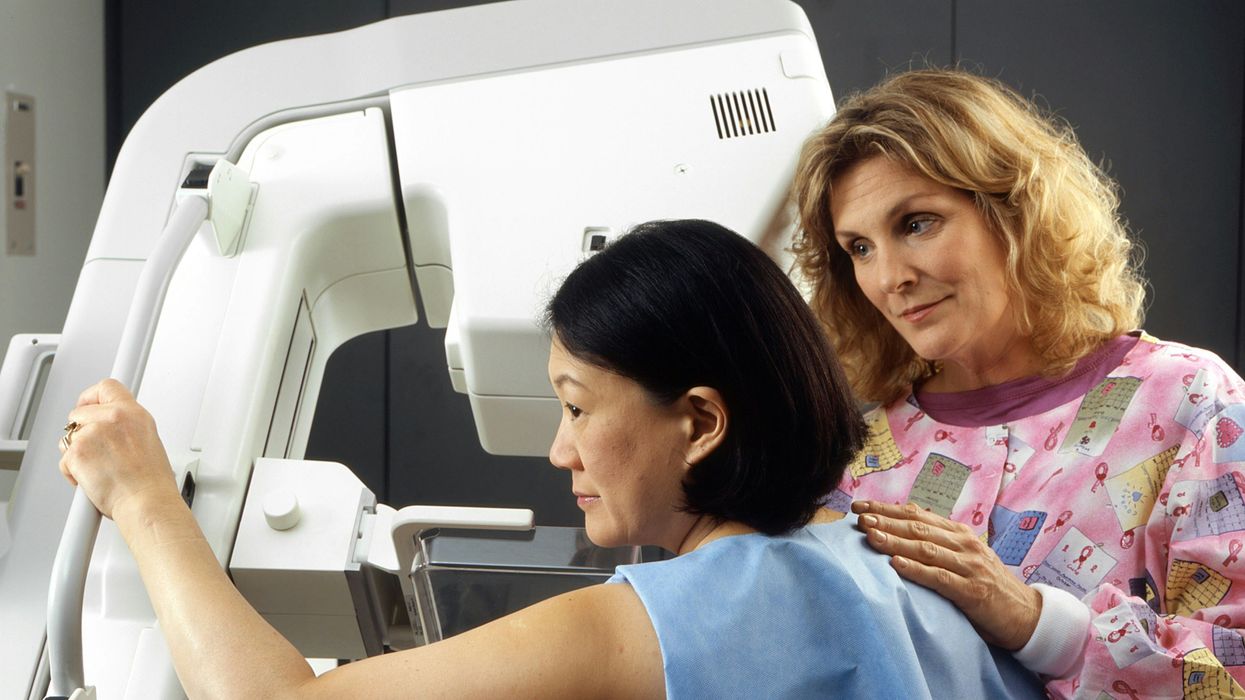
A woman receives a mammogram, which can detect the presence of tumors in a patient's breast.
When a patient is diagnosed with early-stage breast cancer, having surgery to remove the tumor is considered the standard of care. But what happens when a patient can’t have surgery?
Whether it’s due to high blood pressure, advanced age, heart issues, or other reasons, some breast cancer patients don’t qualify for a lumpectomy—one of the most common treatment options for early-stage breast cancer. A lumpectomy surgically removes the tumor while keeping the patient’s breast intact, while a mastectomy removes the entire breast and nearby lymph nodes.
Fortunately, a new technique called cryoablation is now available for breast cancer patients who either aren’t candidates for surgery or don’t feel comfortable undergoing a surgical procedure. With cryoablation, doctors use an ultrasound or CT scan to locate any tumors inside the patient’s breast. They then insert small, needle-like probes into the patient's breast which create an “ice ball” that surrounds the tumor and kills the cancer cells.
Cryoablation has been used for decades to treat cancers of the kidneys and liver—but only in the past few years have doctors been able to use the procedure to treat breast cancer patients. And while clinical trials have shown that cryoablation works for tumors smaller than 1.5 centimeters, a recent clinical trial at Memorial Sloan Kettering Cancer Center in New York has shown that it can work for larger tumors, too.
In this study, doctors performed cryoablation on patients whose tumors were, on average, 2.5 centimeters. The cryoablation procedure lasted for about 30 minutes, and patients were able to go home on the same day following treatment. Doctors then followed up with the patients after 16 months. In the follow-up, doctors found the recurrence rate for tumors after using cryoablation was only 10 percent.
For patients who don’t qualify for surgery, radiation and hormonal therapy is typically used to treat tumors. However, said Yolanda Brice, M.D., an interventional radiologist at Memorial Sloan Kettering Cancer Center, “when treated with only radiation and hormonal therapy, the tumors will eventually return.” Cryotherapy, Brice said, could be a more effective way to treat cancer for patients who can’t have surgery.
“The fact that we only saw a 10 percent recurrence rate in our study is incredibly promising,” she said.