Testing for Any Infectious Disease Could Soon Be As Simple As Peeing On a Stick
In the future, a paper strip reminiscent of a pregnancy test could be used to quickly diagnose the flu and other infectious diseases.
Trying to get a handle on CRISPR news in 2019 can be daunting if you haven't been avidly reading up on it for the last five years.
CRISPR as a diagnostic tool would be a major game changer for medicine and agriculture.
On top of trying to grasp how the science works, and keeping track of its ever expanding applications, you may also have seen coverage of an ongoing legal battle about who owns the intellectual property behind the gene-editing technology CRISPR-Cas9. And then there's the infamous controversy surrounding a scientist who claimed to have used the tool to edit the genomes of two babies in China last year.
But gene editing is not the only application of CRISPR-based biotechnologies. In the future, it may also be used as a tool to diagnose infectious diseases, which could be a major game changer for medicine and agriculture.
How It Works
CRISPR is an acronym for a naturally occurring DNA sequence that normally protects microbes from viruses. It's been compared to a Swiss army knife that can recognize an invader's DNA and precisely destroy it. Repurposed for humans, CRISPR can be paired with a protein called Cas9 that can detect a person's own DNA sequence (usually a problematic one), cut it out, and replace it with a different sequence. Used this way, CRISPR-Cas9 has become a valuable gene-editing tool that is currently being tested to treat numerous genetic diseases, from cancer to blood disorders to blindness.
CRISPR can also be paired with other proteins, like Cas13, which target RNA, the single-stranded twin of DNA that viruses rely on to infect their hosts and cause disease. In a future clinical setting, CRISPR-Cas13 might be used to diagnose whether you have the flu by cutting a target RNA sequence from the virus. That spliced sequence could stick to a paper test strip, causing a band to show up, like on a pregnancy test strip. If the influenza virus and its RNA are not present, no band would show up.
To understand how close to reality this diagnostic scenario is right now, leapsmag chatted with CRISPR pioneer Dr. Feng Zhang, a molecular biologist at the Broad Institute of MIT and Harvard.
What do you think might be the first point of contact that a regular person or patient would have with a CRISPR diagnostic tool?
FZ: I think in the long run it will be great to see this for, say, at-home disease testing, for influenza and other sorts of important public health [concerns]. To be able to get a readout at home, people can potentially quarantine themselves rather than traveling to a hospital and then carrying the risk of spreading that disease to other people as they get to the clinic.
"You could conceivably get a readout during the same office visit, and then the doctor will be able to prescribe the right treatment right away."
Is this just something that people will use at home, or do you also foresee clinical labs at hospitals applying CRISPR-Cas13 to samples that come through?
FZ: I think we'll see applications in both settings, and I think there are advantages to both. One of the nice things about SHERLOCK [a playful acronym for CRISPR-Cas13's longer name, Specific High-sensitivity Enzymatic Reporter unLOCKing] is that it's rapid; you can get a readout fairly quickly. So, right now, what people do in hospitals is they will collect your sample and then they'll send it out to a clinical testing lab, so you wouldn't get a result back until many hours if not several days later. With SHERLOCK, you could conceivably get a readout during the same office visit, and then the doctor will be able to prescribe the right treatment right away.
I just want to clarify that when you say a doctor would take a sample, that's referring to urine, blood, or saliva, correct?
FZ: Right. Yeah, exactly.
Thinking more long term, are there any Holy Grail applications that you hope CRISPR reaches as a diagnostic tool?
FZ: I think in the developed world we'll hopefully see this being used for influenza testing, and many other viral and pathogen-based diseases—both at home and also in the hospital—but I think the even more exciting direction is that this could be used and deployed in parts of the developing world where there isn't a fancy laboratory with elaborate instrumentation. SHERLOCK is relatively inexpensive to develop, and you can turn it into a paper strip test.
Can you quantify what you mean by relatively inexpensive? What range of prices are we talking about here?
FZ: So without accounting for economies of scale, we estimate that it can cost less than a dollar per test. With economy of scale that cost can go even lower.
Is there value in developing what is actually quite an innovative tool in a way that visually doesn't seem innovative because it's reminiscent of a pregnancy test? And I don't mean that as an insult.
FZ: [Laughs] Ultimately, we want the technology to be as accessible as possible, and pregnancy test strips have such a convenient and easy-to-use form. I think modeling after something that people are already familiar with and just changing what's under the hood makes a lot of sense.
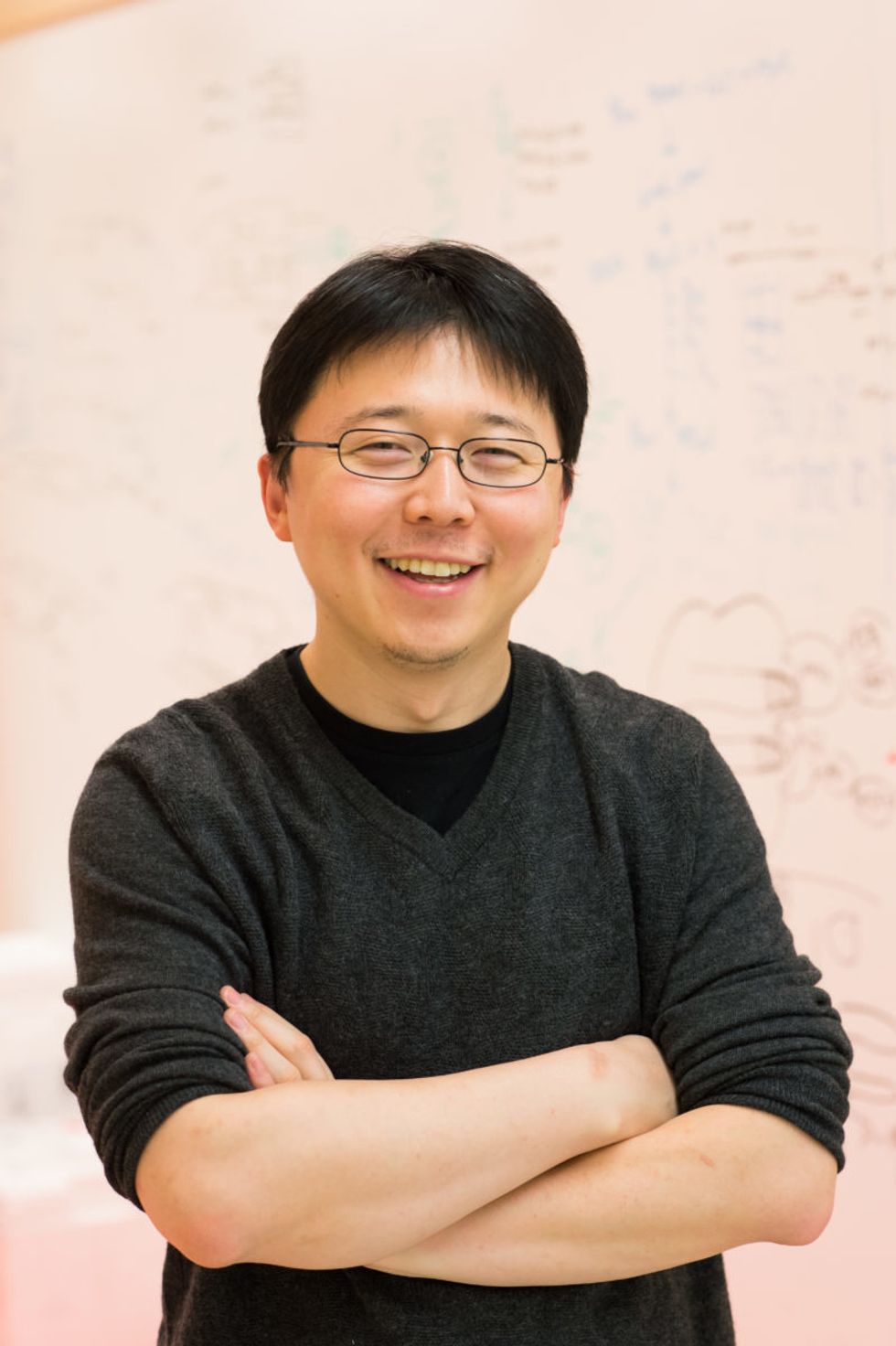

Feng Zhang
(Photo credit: Justin Knight, McGovern Institute)
It's probably one of the most accessible at-home diagnostic tools at this point that people are familiar with.
FZ: Yeah, so if people know how to use that, then using something that's very similar to it should make the option very easy.
You've been quite vocal in calling for some pauses in CRISPR-Cas9 research to make sure it doesn't outpace the ethics of establishing pregnancies with that version of the tool. Do you have any concerns about using CRISPR-Cas13 as a diagnostic tool?
I think overall, the reception for CRISPR-based diagnostics has been overwhelmingly positive. People are very excited about the prospect of using this—for human health and also in agriculture [for] detection of plant infections and plant pathogens, so that farmers will be able to react quickly to infection in the field. If we're looking at contamination of foods by certain bacteria, [food safety] would also be a really exciting application.
Do you feel like the controversies surrounding using CRISPR as a gene-editing tool have overshadowed its potential as a diagnostics tool?
FZ: I don't think so. I think the potential for using CRISPR-Cas9 or CRISPR-Cas12 for gene therapy, and treating disease, has captured people's imaginations, but at the same time, every time I talk with someone about the ability to use CRISPR-Cas13 as a diagnostic tool, people are equally excited. Especially when people see the very simple paper strip that we developed for detecting diseases.
Are CRISPR as a gene-editing tool and CRISPR as a diagnostics tool on different timelines, as far as when the general public might encounter them in their real lives?
FZ: I think they are all moving forward quite quickly. CRISPR as a gene-editing tool is already being deployed in human health and agriculture. We've already seen the approval for the development of growing genome-edited mushrooms, soybeans, and other crop species. So I think people will encounter those in their daily lives in that manner.
Then, of course, for disease treatment, that's progressing rapidly as well. For patients who are affected by sickle cell disease, and also by a degenerative eye disease, clinical trials are already starting in those two areas. Diagnostic tests are also developing quickly, and I think in the coming couple of years, we'll begin to see some of these reaching into the public realm.
"There are probably 7,000 genetic diseases identified today, and most of them don't have any way of being treated."
As far its limits, will it be hard to use CRISPR as a diagnostic tool in situations where we don't necessarily understand the biological underpinnings of a disease?
FZ: CRISPR-Cas13, as a diagnostic tool, at least in the current way that it's implemented, is a detection tool—it's not a discovery tool. So if we don't know what we're looking for, then it's going to be hard to develop Cas13 to detect it. But even in the case of a new infectious disease, if DNA sequencing or RNA sequencing information is available for that new virus, then we can very rapidly program a Cas13-based system to detect it, based on that sequence.
What's something you think the public misunderstands about CRISPR, either in general, or specifically as a diagnostic tool, that you wish were better understood?
FZ: That's a good question. CRISPR-Cas9 and CRISPR-Cas12 as gene editing tools, and also CRISPR-Cas13 as a diagnostic tool, are able to do some things, but there are still a lot of capabilities that need to be further developed. So I think the potential for the technology will unfold over the next decade or so, but it will take some time for the full impact of the technology to really get realized in real life.
What do you think that full impact is?
FZ: There are probably 7,000 genetic diseases identified today, and most of them don't have any way of being treated. It will take some time for CRISPR-Cas9 and Cas12 to be really developed for addressing a larger number of those diseases. And then for CRISPR-based diagnostics, I think you'll see the technology being applied in a couple of initial cases, and it will take some time to develop that more broadly for many other applications.
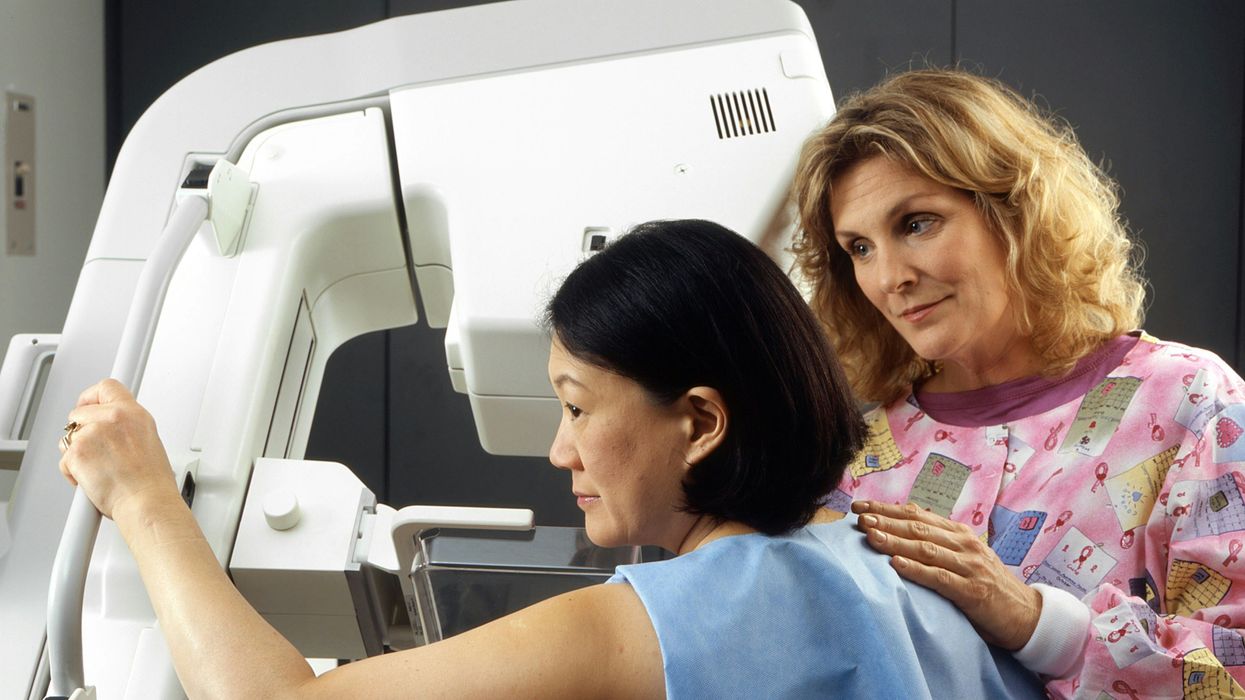
A woman receives a mammogram, which can detect the presence of tumors in a patient's breast.
When a patient is diagnosed with early-stage breast cancer, having surgery to remove the tumor is considered the standard of care. But what happens when a patient can’t have surgery?
Whether it’s due to high blood pressure, advanced age, heart issues, or other reasons, some breast cancer patients don’t qualify for a lumpectomy—one of the most common treatment options for early-stage breast cancer. A lumpectomy surgically removes the tumor while keeping the patient’s breast intact, while a mastectomy removes the entire breast and nearby lymph nodes.
Fortunately, a new technique called cryoablation is now available for breast cancer patients who either aren’t candidates for surgery or don’t feel comfortable undergoing a surgical procedure. With cryoablation, doctors use an ultrasound or CT scan to locate any tumors inside the patient’s breast. They then insert small, needle-like probes into the patient's breast which create an “ice ball” that surrounds the tumor and kills the cancer cells.
Cryoablation has been used for decades to treat cancers of the kidneys and liver—but only in the past few years have doctors been able to use the procedure to treat breast cancer patients. And while clinical trials have shown that cryoablation works for tumors smaller than 1.5 centimeters, a recent clinical trial at Memorial Sloan Kettering Cancer Center in New York has shown that it can work for larger tumors, too.
In this study, doctors performed cryoablation on patients whose tumors were, on average, 2.5 centimeters. The cryoablation procedure lasted for about 30 minutes, and patients were able to go home on the same day following treatment. Doctors then followed up with the patients after 16 months. In the follow-up, doctors found the recurrence rate for tumors after using cryoablation was only 10 percent.
For patients who don’t qualify for surgery, radiation and hormonal therapy is typically used to treat tumors. However, said Yolanda Brice, M.D., an interventional radiologist at Memorial Sloan Kettering Cancer Center, “when treated with only radiation and hormonal therapy, the tumors will eventually return.” Cryotherapy, Brice said, could be a more effective way to treat cancer for patients who can’t have surgery.
“The fact that we only saw a 10 percent recurrence rate in our study is incredibly promising,” she said.
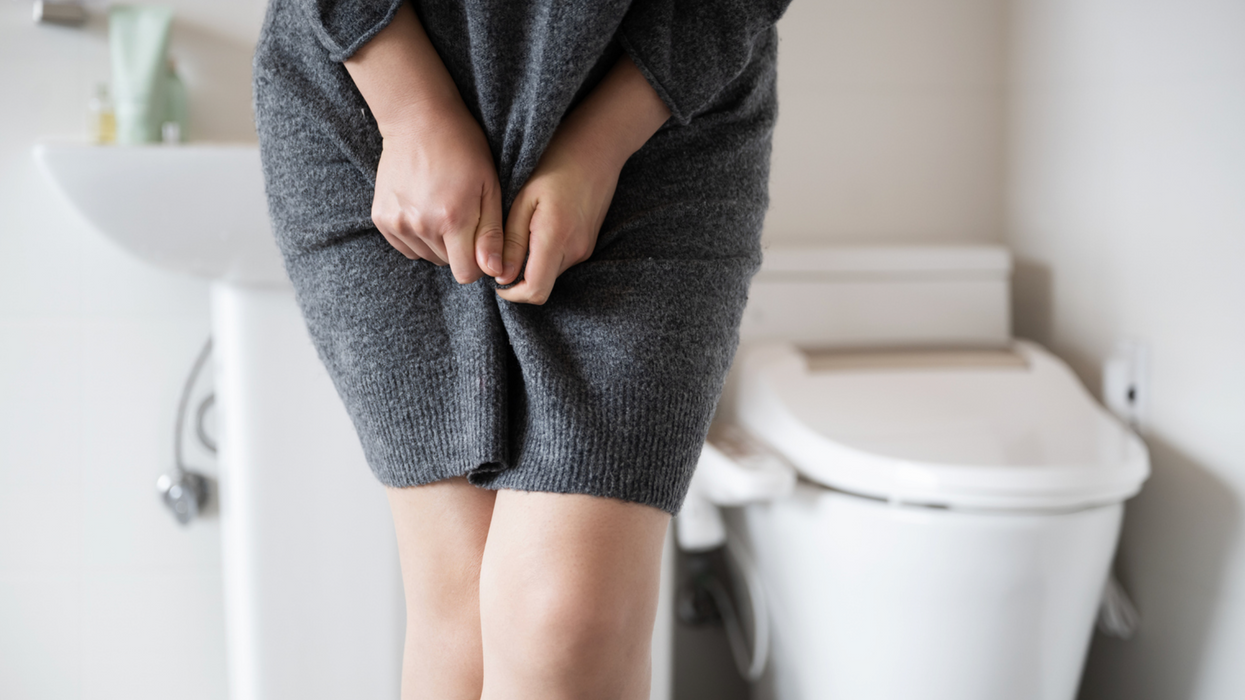
Urinary tract infections account for more than 8 million trips to the doctor each year.
Few things are more painful than a urinary tract infection (UTI). Common in men and women, these infections account for more than 8 million trips to the doctor each year and can cause an array of uncomfortable symptoms, from a burning feeling during urination to fever, vomiting, and chills. For an unlucky few, UTIs can be chronic—meaning that, despite treatment, they just keep coming back.
But new research, presented at the European Association of Urology (EAU) Congress in Paris this week, brings some hope to people who suffer from UTIs.
Clinicians from the Royal Berkshire Hospital presented the results of a long-term, nine-year clinical trial where 89 men and women who suffered from recurrent UTIs were given an oral vaccine called MV140, designed to prevent the infections. Every day for three months, the participants were given two sprays of the vaccine (flavored to taste like pineapple) and then followed over the course of nine years. Clinicians analyzed medical records and asked the study participants about symptoms to check whether any experienced UTIs or had any adverse reactions from taking the vaccine.
The results showed that across nine years, 48 of the participants (about 54%) remained completely infection-free. On average, the study participants remained infection free for 54.7 months—four and a half years.
“While we need to be pragmatic, this vaccine is a potential breakthrough in preventing UTIs and could offer a safe and effective alternative to conventional treatments,” said Gernot Bonita, Professor of Urology at the Alta Bro Medical Centre for Urology in Switzerland, who is also the EAU Chairman of Guidelines on Urological Infections.
The news comes as a relief not only for people who suffer chronic UTIs, but also to doctors who have seen an uptick in antibiotic-resistant UTIs in the past several years. Because UTIs usually require antibiotics, patients run the risk of developing a resistance to the antibiotics, making infections more difficult to treat. A preventative vaccine could mean less infections, less antibiotics, and less drug resistance overall.
“Many of our participants told us that having the vaccine restored their quality of life,” said Dr. Bob Yang, Consultant Urologist at the Royal Berkshire NHS Foundation Trust, who helped lead the research. “While we’re yet to look at the effect of this vaccine in different patient groups, this follow-up data suggests it could be a game-changer for UTI prevention if it’s offered widely, reducing the need for antibiotic treatments.”