Catching colds may help protect kids from Covid
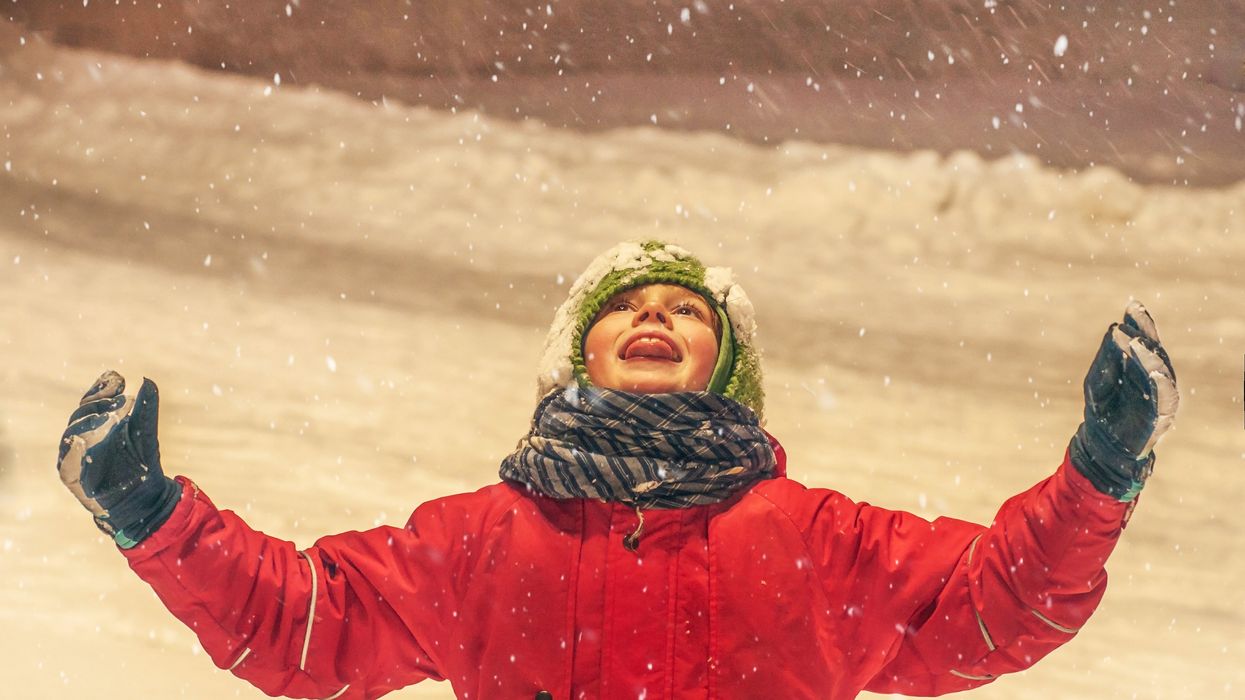

A new study shows that the immune system's response to colds can help prepare it to defend against COVID-19 - but only in the very young.
A common cold virus causes the immune system to produce T cells that also provide protection against SARS-CoV-2, according to new research. The study, published last month in PNAS, shows that this effect is most pronounced in young children. The finding may help explain why most young people who have been exposed to the cold-causing coronavirus have not developed serious cases of COVID-19.
One curiosity stood out in the early days of the COVID-19 pandemic – why were so few kids getting sick. Generally young children and the elderly are the most vulnerable to disease outbreaks, particularly viral infections, either because their immune systems are not fully developed or they are starting to fail.
But solid information on the new infection was so scarce that many public health officials acted on the precautionary principle, assumed a worst-case scenario, and applied the broadest, most restrictive policies to all people to try to contain the coronavirus SARS-CoV-2.
One early thought was that lockdowns worked and kids (ages 6 months to 17 years) simply were not being exposed to the virus. So it was a shock when data started to come in showing that well over half of them carried antibodies to the virus, indicating exposure without getting sick. That trend grew over time and the latest tracking data from the CDC shows that 96.3 percent of kids in the U.S. now carry those antibodies.
Antibodies are relatively quick and easy to measure, but some scientists are exploring whether the reactions of T cells could serve as a more useful measure of immune protection.
But that couldn't be the whole story because antibody protection fades, sometimes as early as a month after exposure and usually within a year. Additionally, SARS-CoV-2 has been spewing out waves of different variants that were more resistant to antibodies generated by their predecessors. The resistance was so significant that over time the FDA withdrew its emergency use authorization for a handful of monoclonal antibodies with earlier approval to treat the infection because they no longer worked.
Antibodies got most of the attention early on because they are part of the first line response of the immune system. Antibodies can bind to viruses and neutralize them, preventing infection. They are relatively quick and easy to measure and even manufacture, but as SARS-CoV-2 showed us, often viruses can quickly evolve to become more resistant to them. Some scientists are exploring whether the reactions of T cells could serve as a more useful measure of immune protection.
Kids, colds and T cells
T cells are part of the immune system that deals with cells once they have become infected. But working with T cells is much more difficult, takes longer, and is more expensive than working with antibodies. So studies often lags behind on this part of the immune system.
A group of researchers led by Annika Karlsson at the Karolinska Institute in Sweden focuses on T cells targeting virus-infected cells and, unsurprisingly, saw that they can play a role in SARS-CoV-2 infection. Other labs have shown that vaccination and natural exposure to the virus generates different patterns of T cell responses.
The Swedes also looked at another member of the coronavirus family, OC43, which circulates widely and is one of several causes of the common cold. The molecular structure of OC43 is similar to its more deadly cousin SARS-CoV-2. Sometimes a T cell response to one virus can produce a cross-reactive response to a similar protein structure in another virus, meaning that T cells will identify and respond to the two viruses in much the same way. Karlsson looked to see if T cells for OC43 from a wide age range of patients were cross-reactive to SARS-CoV-2.
And that is what they found, as reported in the PNAS study last month; there was cross-reactive activity, but it depended on a person’s age. A subset of a certain type of T cells, called mCD4+,, that recognized various protein parts of the cold-causing virus, OC43, expressed on the surface of an infected cell – also recognized those same protein parts from SARS-CoV-2. The T cell response was lower than that generated by natural exposure to SARS-CoV-2, but it was functional and thus could help limit the severity of COVID-19.
“One of the most politicized aspects of our pandemic response was not accepting that children are so much less at risk for severe disease with COVID-19,” because usually young children are among the most vulnerable to pathogens, says Monica Gandhi, professor of medicine at the University of California San Francisco.
“The cross-reactivity peaked at age six when more than half the people tested have a cross-reactive immune response,” says Karlsson, though their sample is too small to say if this finding applies more broadly across the population. The vast majority of children as young as two years had OC43-specific mCD4+ T cell responses. In adulthood, the functionality of both the OC43-specific and the cross-reactive T cells wane significantly, especially with advanced age.
“Considering that the mortality rate in children is the lowest from ages five to nine, and higher in younger children, our results imply that cross-reactive mCD4+ T cells may have a role in the control of SARS-CoV-2 infection in children,” the authors wrote in their paper.
“One of the most politicized aspects of our pandemic response was not accepting that children are so much less at risk for severe disease with COVID-19,” because usually young children are among the most vulnerable to pathogens, says Monica Gandhi, professor of medicine at the University of California San Francisco and author of the book, Endemic: A Post-Pandemic Playbook, to be released by the Mayo Clinic Press this summer. The immune response of kids to SARS-CoV-2 stood our expectations on their head. “We just haven't seen this before, so knowing the mechanism of protection is really important.”
Why the T cell immune response can fade with age is largely unknown. With some viruses such as measles, a single vaccination or infection generates life-long protection. But respiratory tract infections, like SARS-CoV-2, cause a localized infection - specific to certain organs - and that response tends to be shorter lived than systemic infections that affect the entire body. Karlsson suspects the elderly might be exposed to these localized types of viruses less often. Also, frequent continued exposure to a virus that results in reactivation of the memory T cell pool might eventually result in “a kind of immunosenescence or immune exhaustion that is associated with aging,” Karlsson says. https://leaps.org/scientists-just-started-testing-a-new-class-of-drugs-to-slow-and-even-reverse-aging/particle-3 This fading protection is why older people need to be repeatedly vaccinated against SARS-CoV-2.
Policy implications
Following the numbers on COVID-19 infections and severity over the last three years have shown us that healthy young people without risk factors are not likely to develop serious disease. This latest study points to a mechanism that helps explain why. But the inertia of existing policies remains. How should we adjust policy recommendations based on what we know today?
The World Health Organization (WHO) updated their COVID-19 vaccination guidance on March 28. It calls for a focus on vaccinating and boosting those at risk for developing serious disease. The guidance basically shrugged its shoulders when it came to healthy children and young adults receiving vaccinations and boosters against COVID-19. It said the priority should be to administer the “traditional essential vaccines for children,” such as those that protect against measles, rubella, and mumps.
“As an immunologist and a mother, I think that catching a cold or two when you are a kid and otherwise healthy is not that bad for you. Children have a much lower risk of becoming severely ill with SARS-CoV-2,” says Karlsson. She has followed public health guidance in Sweden, which means that her young children have not been vaccinated, but being older, she has received the vaccine and boosters. Gandhi and her children have been vaccinated, but they do not plan on additional boosters.
The WHO got it right in “concentrating on what matters,” which is getting traditional childhood immunizations back on track after their dramatic decline over the last three years, says Gandhi. Nor is there a need for masking in schools, according to a study from the Catalonia region of Spain. It found “no difference in masking and spread in schools,” particularly since tracking data indicate that nearly all young people have been exposed to SARS-CoV-2.
Both researchers lament that public discussion has overemphasized the quickly fading antibody part of the immune response to SARS-CoV-2 compared with the more durable T cell component. They say developing an efficient measure of T cell response for doctors to use in the clinic would help to monitor immunity in people at risk for severe cases of COVID-19 compared with the current method of toting up potential risk factors.
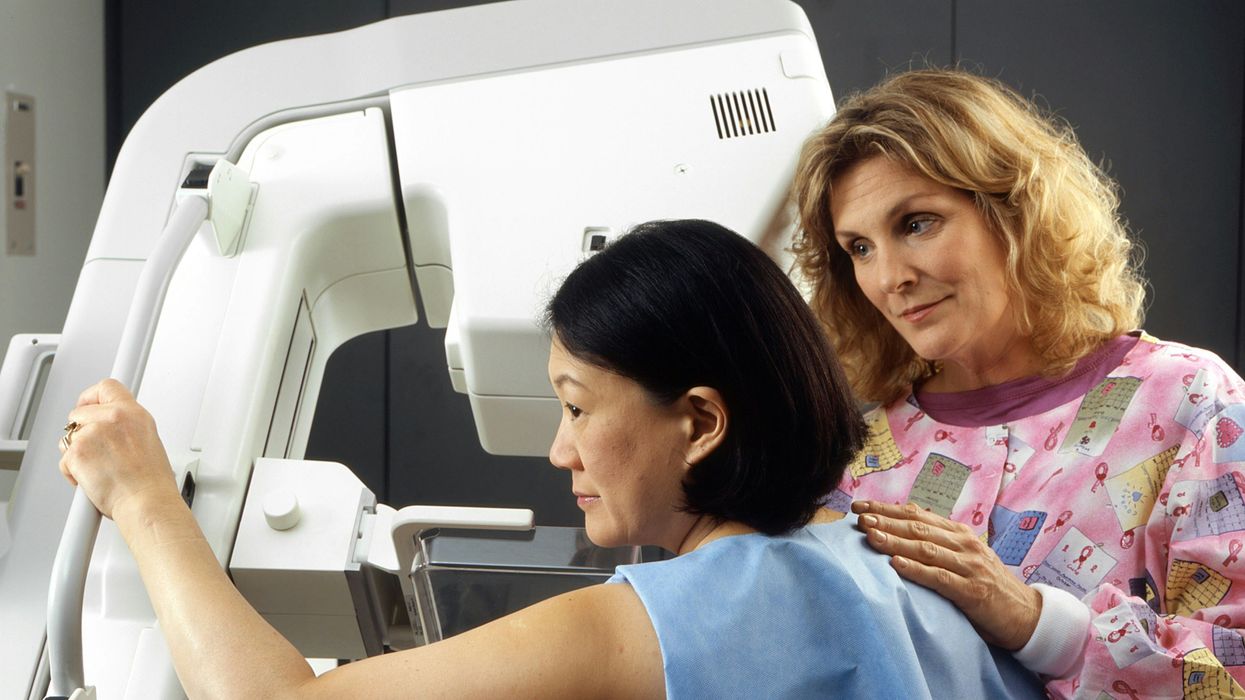
A woman receives a mammogram, which can detect the presence of tumors in a patient's breast.
When a patient is diagnosed with early-stage breast cancer, having surgery to remove the tumor is considered the standard of care. But what happens when a patient can’t have surgery?
Whether it’s due to high blood pressure, advanced age, heart issues, or other reasons, some breast cancer patients don’t qualify for a lumpectomy—one of the most common treatment options for early-stage breast cancer. A lumpectomy surgically removes the tumor while keeping the patient’s breast intact, while a mastectomy removes the entire breast and nearby lymph nodes.
Fortunately, a new technique called cryoablation is now available for breast cancer patients who either aren’t candidates for surgery or don’t feel comfortable undergoing a surgical procedure. With cryoablation, doctors use an ultrasound or CT scan to locate any tumors inside the patient’s breast. They then insert small, needle-like probes into the patient's breast which create an “ice ball” that surrounds the tumor and kills the cancer cells.
Cryoablation has been used for decades to treat cancers of the kidneys and liver—but only in the past few years have doctors been able to use the procedure to treat breast cancer patients. And while clinical trials have shown that cryoablation works for tumors smaller than 1.5 centimeters, a recent clinical trial at Memorial Sloan Kettering Cancer Center in New York has shown that it can work for larger tumors, too.
In this study, doctors performed cryoablation on patients whose tumors were, on average, 2.5 centimeters. The cryoablation procedure lasted for about 30 minutes, and patients were able to go home on the same day following treatment. Doctors then followed up with the patients after 16 months. In the follow-up, doctors found the recurrence rate for tumors after using cryoablation was only 10 percent.
For patients who don’t qualify for surgery, radiation and hormonal therapy is typically used to treat tumors. However, said Yolanda Brice, M.D., an interventional radiologist at Memorial Sloan Kettering Cancer Center, “when treated with only radiation and hormonal therapy, the tumors will eventually return.” Cryotherapy, Brice said, could be a more effective way to treat cancer for patients who can’t have surgery.
“The fact that we only saw a 10 percent recurrence rate in our study is incredibly promising,” she said.
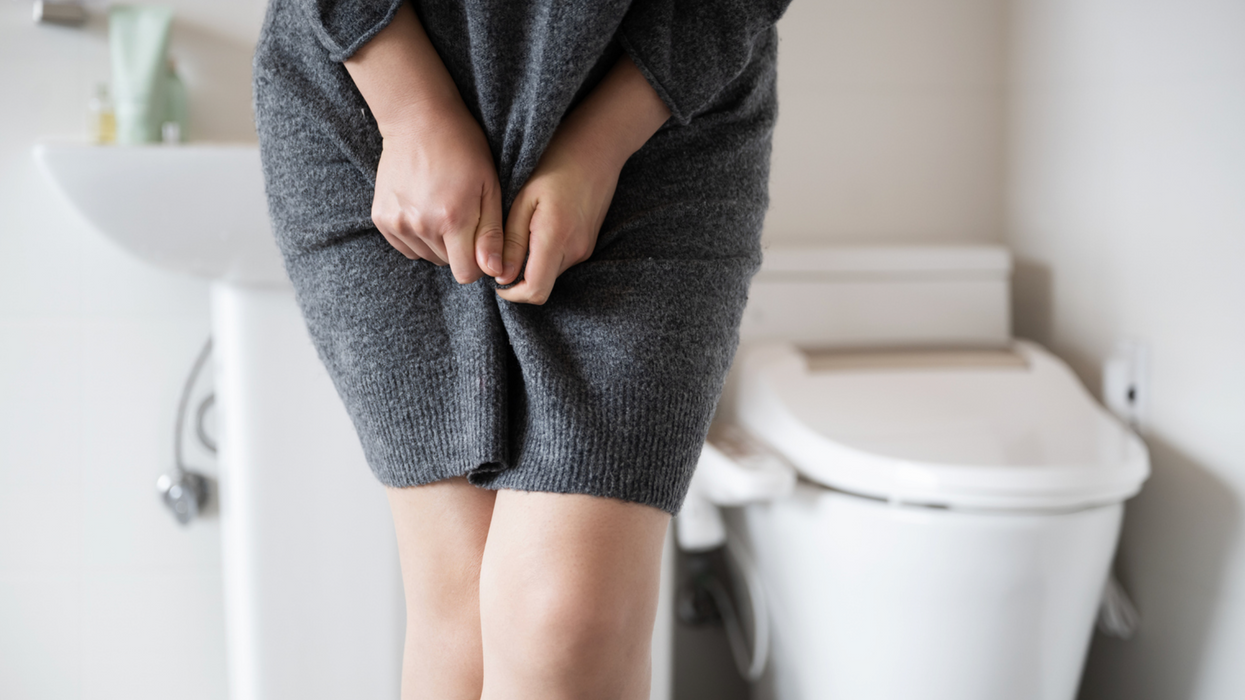
Urinary tract infections account for more than 8 million trips to the doctor each year.
Few things are more painful than a urinary tract infection (UTI). Common in men and women, these infections account for more than 8 million trips to the doctor each year and can cause an array of uncomfortable symptoms, from a burning feeling during urination to fever, vomiting, and chills. For an unlucky few, UTIs can be chronic—meaning that, despite treatment, they just keep coming back.
But new research, presented at the European Association of Urology (EAU) Congress in Paris this week, brings some hope to people who suffer from UTIs.
Clinicians from the Royal Berkshire Hospital presented the results of a long-term, nine-year clinical trial where 89 men and women who suffered from recurrent UTIs were given an oral vaccine called MV140, designed to prevent the infections. Every day for three months, the participants were given two sprays of the vaccine (flavored to taste like pineapple) and then followed over the course of nine years. Clinicians analyzed medical records and asked the study participants about symptoms to check whether any experienced UTIs or had any adverse reactions from taking the vaccine.
The results showed that across nine years, 48 of the participants (about 54%) remained completely infection-free. On average, the study participants remained infection free for 54.7 months—four and a half years.
“While we need to be pragmatic, this vaccine is a potential breakthrough in preventing UTIs and could offer a safe and effective alternative to conventional treatments,” said Gernot Bonita, Professor of Urology at the Alta Bro Medical Centre for Urology in Switzerland, who is also the EAU Chairman of Guidelines on Urological Infections.
The news comes as a relief not only for people who suffer chronic UTIs, but also to doctors who have seen an uptick in antibiotic-resistant UTIs in the past several years. Because UTIs usually require antibiotics, patients run the risk of developing a resistance to the antibiotics, making infections more difficult to treat. A preventative vaccine could mean less infections, less antibiotics, and less drug resistance overall.
“Many of our participants told us that having the vaccine restored their quality of life,” said Dr. Bob Yang, Consultant Urologist at the Royal Berkshire NHS Foundation Trust, who helped lead the research. “While we’re yet to look at the effect of this vaccine in different patient groups, this follow-up data suggests it could be a game-changer for UTI prevention if it’s offered widely, reducing the need for antibiotic treatments.”