Scientists want the salamander's secret: how they regenerate tissue
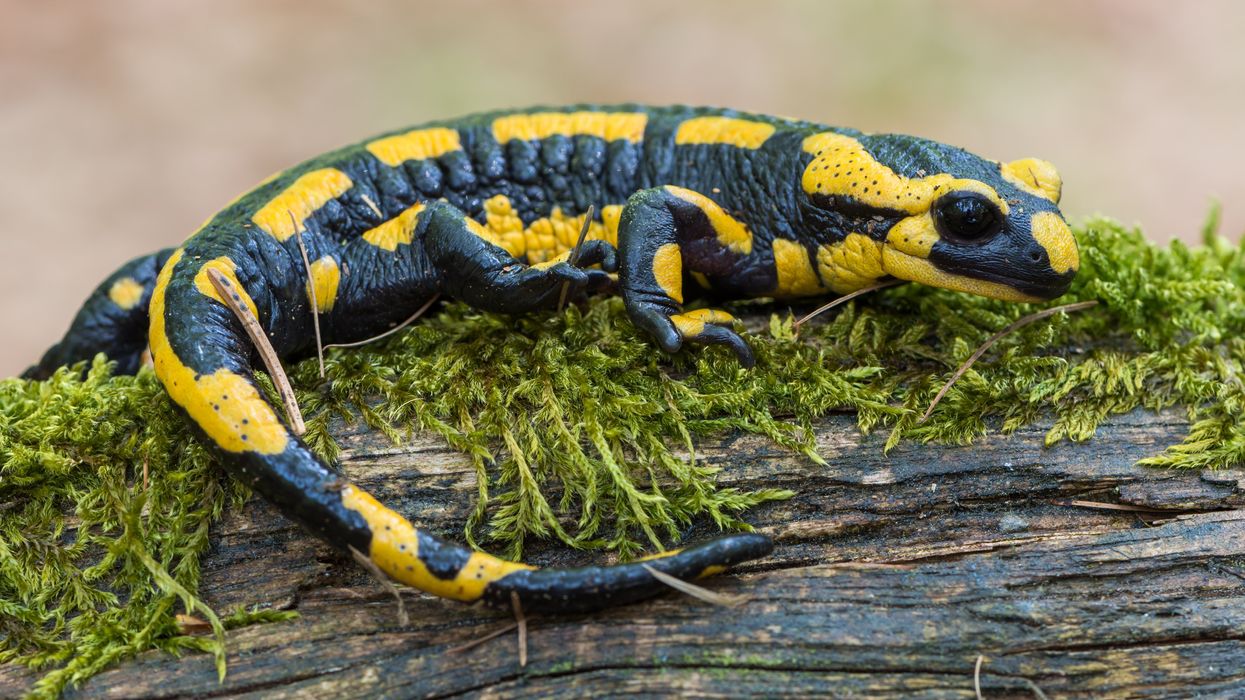

All organisms can repair damaged tissue, but none do it better than salamanders and newts. A surprising area of science could tell us how they manage this feat - and perhaps even help us develop a similar ability.
All organisms have the capacity to repair or regenerate tissue damage. None can do it better than salamanders or newts, which can regenerate an entire severed limb.
That feat has amazed and delighted man from the dawn of time and led to endless attempts to understand how it happens – and whether we can control it for our own purposes. An exciting new clue toward that understanding has come from a surprising source: research on the decline of cells, called cellular senescence.
Senescence is the last stage in the life of a cell. Whereas some cells simply break up or wither and die off, others transition into a zombie-like state where they can no longer divide. In this liminal phase, the cell still pumps out many different molecules that can affect its neighbors and cause low grade inflammation. Senescence is associated with many of the declining biological functions that characterize aging, such as inflammation and genomic instability.
Oddly enough, newts are one of the few species that do not accumulate senescent cells as they age, according to research over several years by Maximina Yun. A research group leader at the Center for Regenerative Therapies Dresden and the Max Planck Institute of Molecular and Cell Biology and Genetics, in Dresden, Germany, Yun discovered that senescent cells were induced at some stages of regeneration of the salamander limb, “and then, as the regeneration progresses, they disappeared, they were eliminated by the immune system,” she says. “They were present at particular times and then they disappeared.”
Senescent cells added to the edges of the wound helped the healthy muscle cells to “dedifferentiate,” essentially turning back the developmental clock of those cells into more primitive states.
Previous research on senescence in aging had suggested, logically enough, that applying those cells to the stump of a newly severed salamander limb would slow or even stop its regeneration. But Yun stood that idea on its head. She theorized that senescent cells might also play a role in newt limb regeneration, and she tested it by both adding and removing senescent cells from her animals. It turned out she was right, as the newt limbs grew back faster than normal when more senescent cells were included.
Senescent cells added to the edges of the wound helped the healthy muscle cells to “dedifferentiate,” essentially turning back the developmental clock of those cells into more primitive states, which could then be turned into progenitors, a cell type in between stem cells and specialized cells, needed to regrow the muscle tissue of the missing limb. “We think that this ability to dedifferentiate is intrinsically a big part of why salamanders can regenerate all these very complex structures, which other organisms cannot,” she explains.
Yun sees regeneration as a two part problem. First, the cells must be able to sense that their neighbors from the lost limb are not there anymore. Second, they need to be able to produce the intermediary progenitors for regeneration, , to form what is missing. “Molecularly, that must be encoded like a 3D map,” she says, otherwise the new tissue might grow back as a blob, or liver, or fin instead of a limb.
Wound healing
Another recent study, this time at the Mayo Clinic, provides evidence supporting the role of senescent cells in regeneration. Looking closely at molecules that send information between cells in the wound of a mouse, the researchers found that senescent cells appeared near the start of the healing process and then disappeared as healing progressed. In contrast, persistent senescent cells were the hallmark of a chronic wound that did not heal properly. The function and significance of senescence cells depended on both the timing and the context of their environment.
The paper suggests that senescent cells are not all the same. That has become clearer as researchers have been able to identify protein markers on the surface of some senescent cells. The patterns of these proteins differ for some senescent cells compared to others. In biology, such physical differences suggest functional differences, so it is becoming increasingly likely there are subsets of senescent cells with differing functions that have not yet been identified.
There are disagreements within the research community as to whether newts have acquired their regenerative capacity through a unique evolutionary change, or if other animals, including humans, retain this capacity buried somewhere in their genes.
Scientists initially thought that senescent cells couldn’t play a role in regeneration because they could no longer reproduce, says Anthony Atala, a practicing surgeon and bioengineer who leads the Wake Forest Institute for Regenerative Medicine in North Carolina. But Yun’s study points in the other direction. “What this paper shows clearly is that these cells have the potential to be involved in tissue regeneration [in newts]. The question becomes, will these cells be able to do the same in humans.”
As our knowledge of senescent cells increases, Atala thinks we need to embrace a new analogy to help understand them: humans in retirement. They “have acquired a lot of wisdom throughout their whole life and they can help younger people and mentor them to grow to their full potential. We're seeing the same thing with these cells,” he says. They are no longer putting energy into their own reproduction, but the signaling molecules they secrete “can help other cells around them to regenerate.”
There are disagreements within the research community as to whether newts have acquired their regenerative capacity through a unique evolutionary change, or if other animals, including humans, retain this capacity buried somewhere in their genes. If so, it seems that our genes are unable to express this ability, perhaps as part of a tradeoff in acquiring other traits. It is a fertile area of research.
Dedifferentiation is likely to become an important process in the field of regenerative medicine. One extreme example: a lab has been able to turn back the clock and reprogram adult male skin cells into female eggs, a potential milestone in reproductive health. It will be more difficult to control just how far back one wishes to go in the cell's dedifferentiation – part way or all the way back into a stem cell – and then direct it down a different developmental pathway. Yun is optimistic we can learn these tricks from newts.
Senolytics
A growing field of research is using drugs called senolytics to remove senescent cells and slow or even reverse disease of aging.
“Senolytics are great, but senolytics target different types of senescence,” Yun says. “If senescent cells have positive effects in the context of regeneration, of wound healing, then maybe at the beginning of the regeneration process, you may not want to take them out for a little while.”
“If you look at pretty much all biological systems, too little or too much of something can be bad, you have to be in that central zone” and at the proper time, says Atala. “That's true for proteins, sugars, and the drugs that you take. I think the same thing is true for these cells. Why would they be different?”
Our growing understanding that senescence is not a single thing but a variety of things likely means that effective senolytic drugs will not resemble a single sledge hammer but more a carefully manipulated scalpel where some types of senescent cells are removed while others are added. Combinations and timing could be crucial, meaning the difference between regenerating healthy tissue, a scar, or worse.
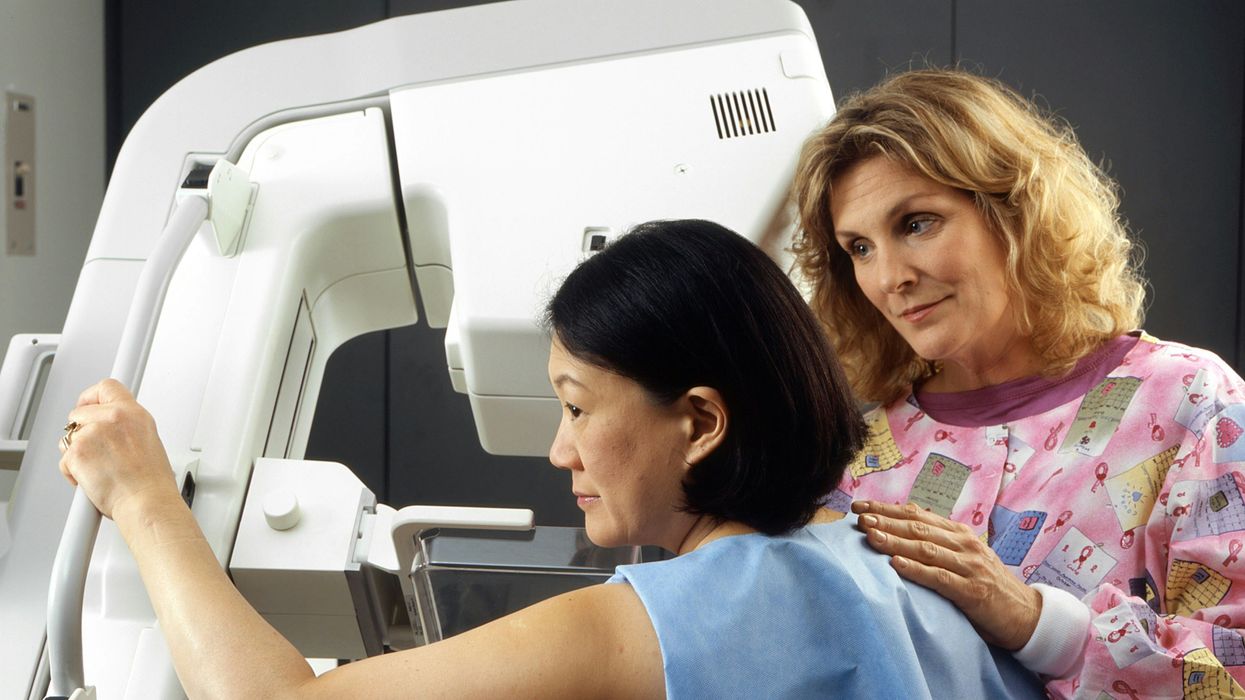
A woman receives a mammogram, which can detect the presence of tumors in a patient's breast.
When a patient is diagnosed with early-stage breast cancer, having surgery to remove the tumor is considered the standard of care. But what happens when a patient can’t have surgery?
Whether it’s due to high blood pressure, advanced age, heart issues, or other reasons, some breast cancer patients don’t qualify for a lumpectomy—one of the most common treatment options for early-stage breast cancer. A lumpectomy surgically removes the tumor while keeping the patient’s breast intact, while a mastectomy removes the entire breast and nearby lymph nodes.
Fortunately, a new technique called cryoablation is now available for breast cancer patients who either aren’t candidates for surgery or don’t feel comfortable undergoing a surgical procedure. With cryoablation, doctors use an ultrasound or CT scan to locate any tumors inside the patient’s breast. They then insert small, needle-like probes into the patient's breast which create an “ice ball” that surrounds the tumor and kills the cancer cells.
Cryoablation has been used for decades to treat cancers of the kidneys and liver—but only in the past few years have doctors been able to use the procedure to treat breast cancer patients. And while clinical trials have shown that cryoablation works for tumors smaller than 1.5 centimeters, a recent clinical trial at Memorial Sloan Kettering Cancer Center in New York has shown that it can work for larger tumors, too.
In this study, doctors performed cryoablation on patients whose tumors were, on average, 2.5 centimeters. The cryoablation procedure lasted for about 30 minutes, and patients were able to go home on the same day following treatment. Doctors then followed up with the patients after 16 months. In the follow-up, doctors found the recurrence rate for tumors after using cryoablation was only 10 percent.
For patients who don’t qualify for surgery, radiation and hormonal therapy is typically used to treat tumors. However, said Yolanda Brice, M.D., an interventional radiologist at Memorial Sloan Kettering Cancer Center, “when treated with only radiation and hormonal therapy, the tumors will eventually return.” Cryotherapy, Brice said, could be a more effective way to treat cancer for patients who can’t have surgery.
“The fact that we only saw a 10 percent recurrence rate in our study is incredibly promising,” she said.
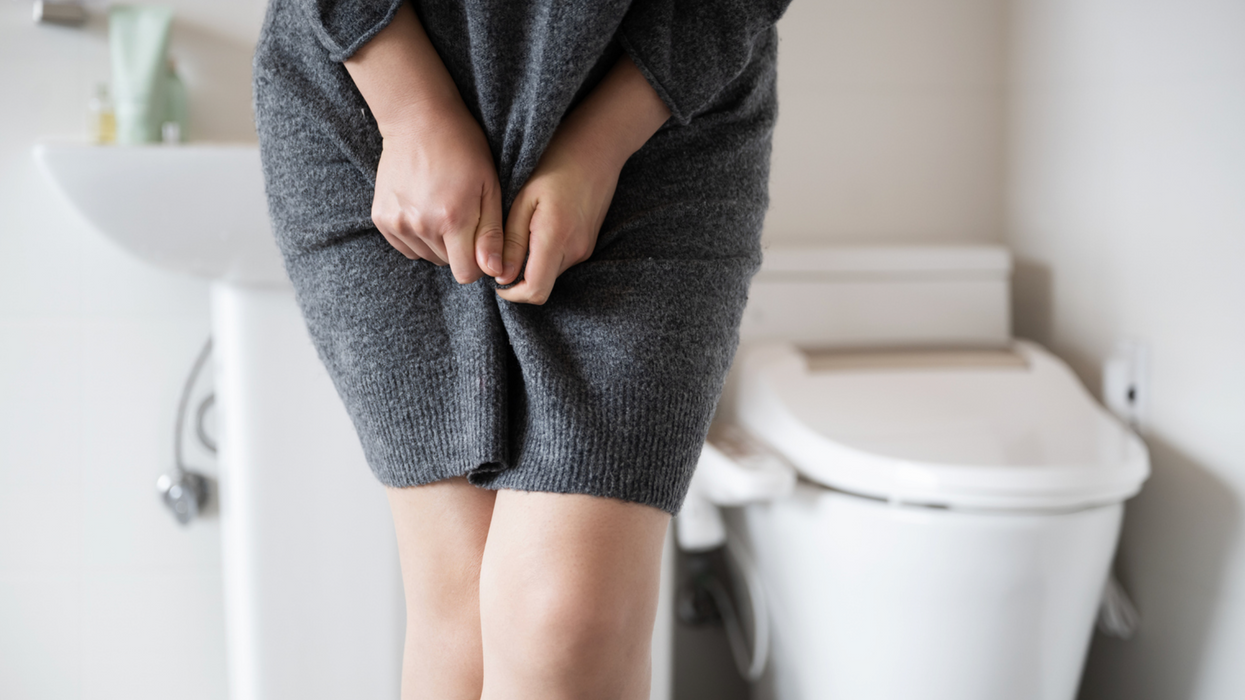
Urinary tract infections account for more than 8 million trips to the doctor each year.
Few things are more painful than a urinary tract infection (UTI). Common in men and women, these infections account for more than 8 million trips to the doctor each year and can cause an array of uncomfortable symptoms, from a burning feeling during urination to fever, vomiting, and chills. For an unlucky few, UTIs can be chronic—meaning that, despite treatment, they just keep coming back.
But new research, presented at the European Association of Urology (EAU) Congress in Paris this week, brings some hope to people who suffer from UTIs.
Clinicians from the Royal Berkshire Hospital presented the results of a long-term, nine-year clinical trial where 89 men and women who suffered from recurrent UTIs were given an oral vaccine called MV140, designed to prevent the infections. Every day for three months, the participants were given two sprays of the vaccine (flavored to taste like pineapple) and then followed over the course of nine years. Clinicians analyzed medical records and asked the study participants about symptoms to check whether any experienced UTIs or had any adverse reactions from taking the vaccine.
The results showed that across nine years, 48 of the participants (about 54%) remained completely infection-free. On average, the study participants remained infection free for 54.7 months—four and a half years.
“While we need to be pragmatic, this vaccine is a potential breakthrough in preventing UTIs and could offer a safe and effective alternative to conventional treatments,” said Gernot Bonita, Professor of Urology at the Alta Bro Medical Centre for Urology in Switzerland, who is also the EAU Chairman of Guidelines on Urological Infections.
The news comes as a relief not only for people who suffer chronic UTIs, but also to doctors who have seen an uptick in antibiotic-resistant UTIs in the past several years. Because UTIs usually require antibiotics, patients run the risk of developing a resistance to the antibiotics, making infections more difficult to treat. A preventative vaccine could mean less infections, less antibiotics, and less drug resistance overall.
“Many of our participants told us that having the vaccine restored their quality of life,” said Dr. Bob Yang, Consultant Urologist at the Royal Berkshire NHS Foundation Trust, who helped lead the research. “While we’re yet to look at the effect of this vaccine in different patient groups, this follow-up data suggests it could be a game-changer for UTI prevention if it’s offered widely, reducing the need for antibiotic treatments.”