How Leqembi became the biggest news in Alzheimer’s disease in 40 years, and what comes next
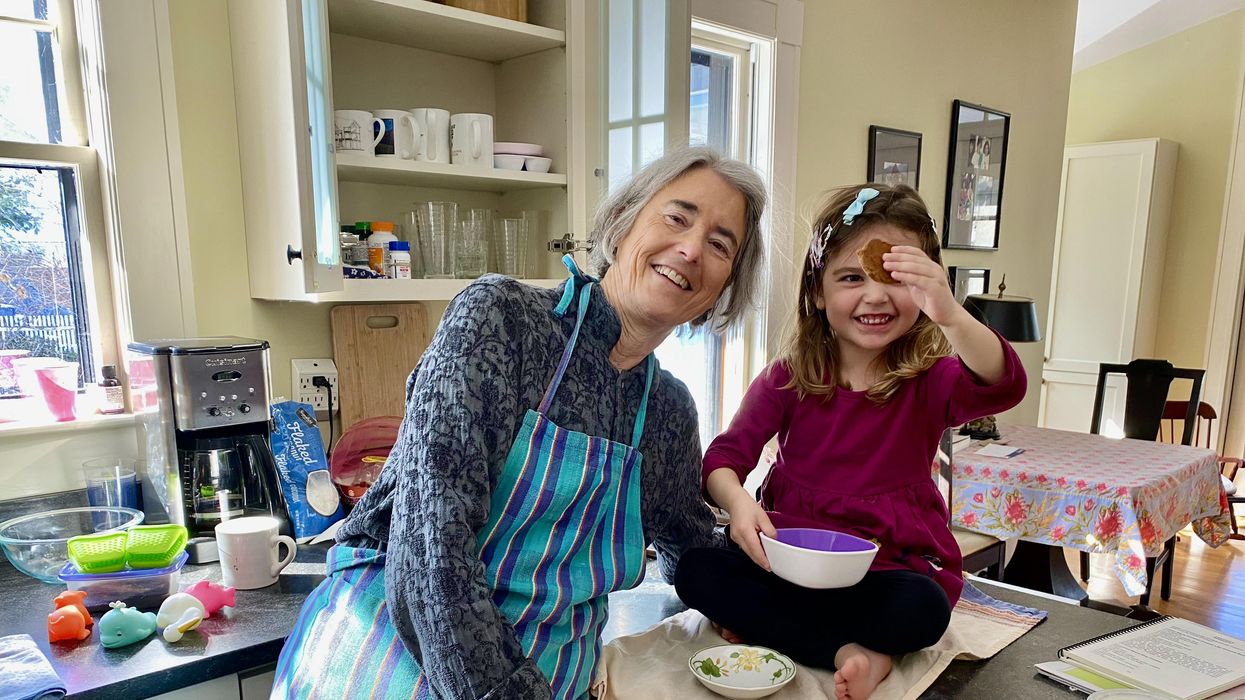

Betsy Groves, 73, with her granddaughter. Groves learned in 2021 that she has Alzheimer's disease. She hopes to take Leqembi, a drug approved by the FDA last week.
A few months ago, Betsy Groves traveled less than a mile from her home in Cambridge, Mass. to give a talk to a bunch of scientists. The scientists, who worked for the pharmaceutical companies Biogen and Eisai, wanted to know how she lived her life, how she thought about her future, and what it was like when a doctor’s appointment in 2021 gave her the worst possible news. Groves, 73, has Alzheimer’s disease. She caught it early, through a lumbar puncture that showed evidence of amyloid, an Alzheimer’s hallmark, in her cerebrospinal fluid. As a way of dealing with her diagnosis, she joined the Alzheimer’s Association’s National Early-Stage Advisory Board, which helped her shift into seeing her diagnosis as something she could use to help others.
After her talk, Groves stayed for lunch with the scientists, who were eager to put a face to their work. Biogen and Eisai were about to release the first drug to successfully combat Alzheimer’s in 40 years of experimental disaster. Their drug, which is known by the scientific name lecanemab and the marketing name Leqembi, was granted accelerated approval by the U.S. Food and Drug Administration last Friday, Jan. 6, after a study in 1,800 people showed that it reduced cognitive decline by 27 percent over 18 months.
It is no exaggeration to say that this result is a huge deal. The field of Alzheimer’s drug development has been absolutely littered with failures. Almost everything researchers have tried has tanked in clinical trials. “Most of the things that we've done have proven not to be effective, and it's not because we haven’t been taking a ton of shots at goal,” says Anton Porsteinsson, director of the University of Rochester Alzheimer's Disease Care, Research, and Education Program, who worked on the lecanemab trial. “I think it's fair to say you don't survive in this field unless you're an eternal optimist.”
As far back as 1984, a cure looked like it was within reach: Scientists discovered that the sticky plaques that develop in the brains of those who have Alzheimer’s are made up of a protein fragment called beta-amyloid. Buildup of beta-amyloid seemed to be sufficient to disrupt communication between, and eventually kill, memory cells. If that was true, then the cure should be straightforward: Stop the buildup of beta-amyloid; stop the Alzheimer’s disease.
It wasn’t so simple. Over the next 38 years, hundreds of drugs designed either to interfere with the production of abnormal amyloid or to clear it from the brain flamed out in trials. It got so bad that neuroscience drug divisions at major pharmaceutical companies (AstraZeneca, Pfizer, Bristol-Myers, GSK, Amgen) closed one by one, leaving the field to smaller, scrappier companies, like Cambridge-based Biogen and Tokyo-based Eisai. Some scientists began to dismiss the amyloid hypothesis altogether: If this protein fragment was so important to the disease, why didn’t ridding the brain of it do anything for patients? There was another abnormal protein that showed up in the brains of Alzheimer’s patients, called tau. Some researchers defected to the tau camp, or came to believe the proteins caused damage in combination.
The situation came to a head in 2021, when the FDA granted provisional approval to a drug called aducanumab, marketed as Aduhelm, against the advice of its own advisory council. The approval was based on proof that Aduhelm reduced beta-amyloid in the brain, even though one research trial showed it had no effect on people’s symptoms or daily life. Aduhelm could also cause serious side effects, like brain swelling and amyloid related imaging abnormalities (known as ARIA, these are basically micro-bleeds that appear on MRI scans). Without a clear benefit to memory loss that would make these risks worth it, Medicare refused to pay for Aduhelm among the general population. Two congressional committees launched an investigation into the drug’s approval, citing corporate greed, lapses in protocol, and an unjustifiably high price. (Aduhelm was also produced by the pharmaceutical company Biogen.)
To be clear, Leqembi is not the cure Alzheimer’s researchers hope for. While the drug is the first to show clear signs of a clinical benefit, the scientific establishment is split on how much of a difference Leqembi will make in the real world.
So far, Leqembi is like Aduhelm in that it has been given accelerated approval only for its ability to remove amyloid from the brain. Both are monoclonal antibodies that direct the immune system to attack and clear dysfunctional beta-amyloid. The difference is that, while that’s all Aduhelm was ever shown to do, Leqembi’s makers have already asked the FDA to give it full approval – a decision that would increase the likelihood that Medicare will cover it – based on data that show it also improves Alzheimer’s sufferer’s lives. Leqembi targets a different type of amyloid, a soluble version called “protofibrils,” and that appears to change the effect. “It can give individuals and their families three, six months longer to be participating in daily life and living independently,” says Claire Sexton, PhD, senior director of scientific programs & outreach for the Alzheimer's Association. “These types of changes matter for individuals and for their families.”
To be clear, Leqembi is not the cure Alzheimer’s researchers hope for. It does not halt or reverse the disease, and people do not get better. While the drug is the first to show clear signs of a clinical benefit, the scientific establishment is split on how much of a difference Leqembi will make in the real world. It has “a rather small effect,” wrote NIH Alzheimer’s researcher Madhav Thambisetty, MD, PhD, in an email to Leaps.org. “It is unclear how meaningful this difference will be to patients, and it is unlikely that this level of difference will be obvious to a patient (or their caregivers).” Another issue is cost: Leqembi will become available to patients later this month, but Eisai is setting the price at $26,500 per year, meaning that very few patients will be able to afford it unless Medicare chooses to reimburse them for it.
The same side effects that plagued Aduhelm are common in Leqembi treatment as well. In many patients, amyloid doesn’t just accumulate around neurons, it also forms deposits in the walls of blood vessels. Blood vessels that are shot through with amyloid are more brittle. If you infuse a drug that targets amyloid, brittle blood vessels in the brain can develop leakage that results in swelling or bleeds. Most of these come with no symptoms, and are only seen during testing, which is why they are called “imaging abnormalities.” But in situations where patients have multiple diseases or are prescribed incompatible drugs, they can be serious enough to cause death. The three deaths reported from Leqembi treatment (so far) are enough to make Thambisetty wonder “how well the drug may be tolerated in real world clinical practice where patients are likely to be sicker and have multiple other medical conditions in contrast to carefully selected patients in clinical trials.”
Porsteinsson believes that earlier detection of Alzheimer’s disease will be the next great advance in treatment, a more important step forward than Leqembi’s approval.
Still, there are reasons to be excited. A successful Alzheimer’s drug can pave the way for combination studies, in which patients try a known effective drug alongside newer, more experimental ones; or preventative studies, which take place years before symptoms occur. It also represents enormous strides in researchers’ understanding of the disease. For example, drug dosages have increased massively—in some cases quadrupling—from the early days of Alzheimer’s research. And patient selection for studies has changed drastically as well. Doctors now know that you’ve got to catch the disease early, through PET-scans or CSF tests for amyloid, if you want any chance of changing its course.
Porsteinsson believes that earlier detection of Alzheimer’s disease will be the next great advance in treatment, a more important step forward than Leqembi’s approval. His lab already uses blood tests for different types of amyloid, for different types of tau, and for measures of neuroinflammation, neural damage, and synaptic health, but commercially available versions from companies like C2N, Quest, and Fuji Rebio are likely to hit the market in the next couple of years. “[They are] going to transform the diagnosis of Alzheimer's disease,” Porsteinsson says. “If someone is experiencing memory problems, their physicians will be able to order a blood test that will tell us if this is the result of changes in your brain due to Alzheimer's disease. It will ultimately make it much easier to identify people at a very early stage of the disease, where they are most likely to benefit from treatment.”
Learn more about new blood tests to detect Alzheimer's
Early detection can help patients for more philosophical reasons as well. Betsy Groves credits finding her Alzheimer’s early with giving her the space to understand and process the changes that were happening to her before they got so bad that she couldn’t. She has been able to update her legal documents and, through her role on the Advisory Group, help the Alzheimer’s Association with developing its programs and support services for people in the early stages of the disease. She still drives, and because she and her husband love to travel, they are hoping to get out of grey, rainy Cambridge and off to Texas or Arizona this spring.
Because her Alzheimer’s disease involves amyloid deposits (a “substantial portion” do not, says Claire Sexton, which is an additional complication for research), and has not yet reached an advanced stage, Groves may be a good candidate to try Leqembi. She says she’d welcome the opportunity to take it. If she can get access, Groves hopes the drug will give her more days to be fully functioning with her husband, daughters, and three grandchildren. Mostly, she avoids thinking about what the latter stages of Alzheimer’s might be like, but she knows the time will come when it will be her reality. “So whatever lecanemab can do to extend my more productive ways of engaging with relationships in the world,” she says. “I'll take that in a minute.”
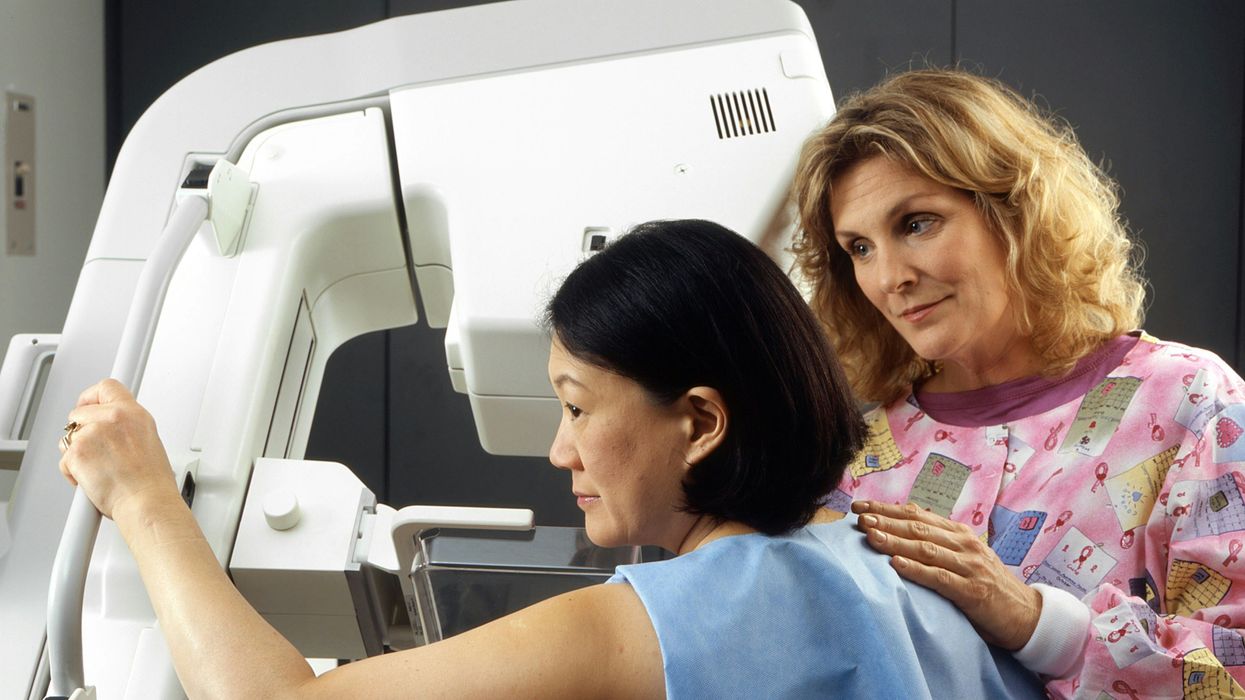
A woman receives a mammogram, which can detect the presence of tumors in a patient's breast.
When a patient is diagnosed with early-stage breast cancer, having surgery to remove the tumor is considered the standard of care. But what happens when a patient can’t have surgery?
Whether it’s due to high blood pressure, advanced age, heart issues, or other reasons, some breast cancer patients don’t qualify for a lumpectomy—one of the most common treatment options for early-stage breast cancer. A lumpectomy surgically removes the tumor while keeping the patient’s breast intact, while a mastectomy removes the entire breast and nearby lymph nodes.
Fortunately, a new technique called cryoablation is now available for breast cancer patients who either aren’t candidates for surgery or don’t feel comfortable undergoing a surgical procedure. With cryoablation, doctors use an ultrasound or CT scan to locate any tumors inside the patient’s breast. They then insert small, needle-like probes into the patient's breast which create an “ice ball” that surrounds the tumor and kills the cancer cells.
Cryoablation has been used for decades to treat cancers of the kidneys and liver—but only in the past few years have doctors been able to use the procedure to treat breast cancer patients. And while clinical trials have shown that cryoablation works for tumors smaller than 1.5 centimeters, a recent clinical trial at Memorial Sloan Kettering Cancer Center in New York has shown that it can work for larger tumors, too.
In this study, doctors performed cryoablation on patients whose tumors were, on average, 2.5 centimeters. The cryoablation procedure lasted for about 30 minutes, and patients were able to go home on the same day following treatment. Doctors then followed up with the patients after 16 months. In the follow-up, doctors found the recurrence rate for tumors after using cryoablation was only 10 percent.
For patients who don’t qualify for surgery, radiation and hormonal therapy is typically used to treat tumors. However, said Yolanda Brice, M.D., an interventional radiologist at Memorial Sloan Kettering Cancer Center, “when treated with only radiation and hormonal therapy, the tumors will eventually return.” Cryotherapy, Brice said, could be a more effective way to treat cancer for patients who can’t have surgery.
“The fact that we only saw a 10 percent recurrence rate in our study is incredibly promising,” she said.
Urinary tract infections account for more than 8 million trips to the doctor each year.
Few things are more painful than a urinary tract infection (UTI). Common in men and women, these infections account for more than 8 million trips to the doctor each year and can cause an array of uncomfortable symptoms, from a burning feeling during urination to fever, vomiting, and chills. For an unlucky few, UTIs can be chronic—meaning that, despite treatment, they just keep coming back.
But new research, presented at the European Association of Urology (EAU) Congress in Paris this week, brings some hope to people who suffer from UTIs.
Clinicians from the Royal Berkshire Hospital presented the results of a long-term, nine-year clinical trial where 89 men and women who suffered from recurrent UTIs were given an oral vaccine called MV140, designed to prevent the infections. Every day for three months, the participants were given two sprays of the vaccine (flavored to taste like pineapple) and then followed over the course of nine years. Clinicians analyzed medical records and asked the study participants about symptoms to check whether any experienced UTIs or had any adverse reactions from taking the vaccine.
The results showed that across nine years, 48 of the participants (about 54%) remained completely infection-free. On average, the study participants remained infection free for 54.7 months—four and a half years.
“While we need to be pragmatic, this vaccine is a potential breakthrough in preventing UTIs and could offer a safe and effective alternative to conventional treatments,” said Gernot Bonita, Professor of Urology at the Alta Bro Medical Centre for Urology in Switzerland, who is also the EAU Chairman of Guidelines on Urological Infections.
The news comes as a relief not only for people who suffer chronic UTIs, but also to doctors who have seen an uptick in antibiotic-resistant UTIs in the past several years. Because UTIs usually require antibiotics, patients run the risk of developing a resistance to the antibiotics, making infections more difficult to treat. A preventative vaccine could mean less infections, less antibiotics, and less drug resistance overall.
“Many of our participants told us that having the vaccine restored their quality of life,” said Dr. Bob Yang, Consultant Urologist at the Royal Berkshire NHS Foundation Trust, who helped lead the research. “While we’re yet to look at the effect of this vaccine in different patient groups, this follow-up data suggests it could be a game-changer for UTI prevention if it’s offered widely, reducing the need for antibiotic treatments.”