3 Futuristic Biotech Programs the U.S. Government Is Funding Right Now
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.
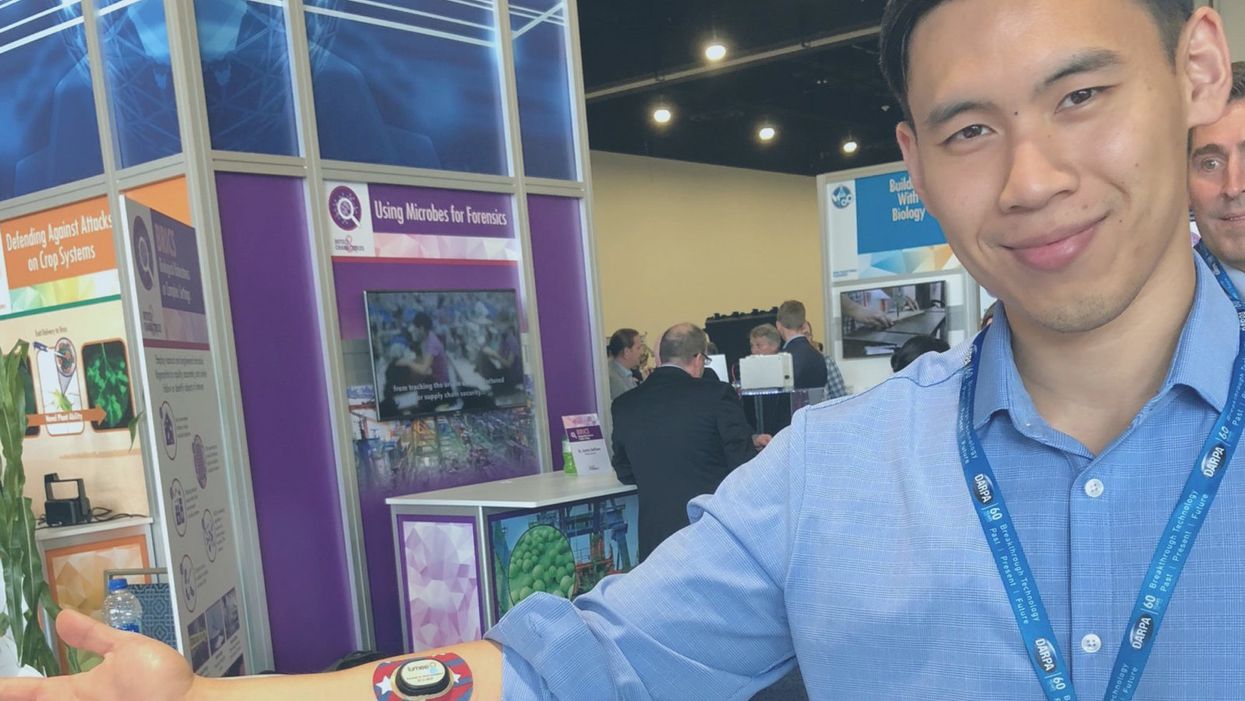

Biomedical engineer Kevin Zhao has a sensor in his arm and his chest that monitors his oxygen level in those tissues in real time.
Last month, at a conference celebrating DARPA, the research arm of the Defense Department, FBI Special Agent Edward You declared, "The 21st century will be the revolution of the life sciences."
Biomedical engineer Kevin Zhao has a sensor in his arm and chest that monitors his oxygen level in real time.
Indeed, four years ago, the agency dedicated a new office solely to advancing biotechnology. Its primary goal is to combat bioterrorism, protect U.S. forces, and promote warfighter readiness. But its research could also carry over to improve health care for the general public.
With an annual budget of about $3 billion, DARPA's employees oversee about 250 research and development programs, working with contractors from corporations, universities, and government labs to bring new technologies to life.
Check out these three current programs:
1) IMPLANTABLE SENSORS TO MEASURE OXYGEN, LACTATE, AND GLUCOSE LEVELS IN REAL TIME
Biomedical engineer Kevin Zhao has a sensor in his arm and his chest that monitors his oxygen level in those tissues in real time. With funding from DARPA for the program "In Vivo Nanoplatforms," he developed soft, flexible hydrogels that are injected just beneath the skin to perform the monitoring and that sync to a smartphone app to give the user immediate health insights.
A first-in-man trial for the glucose sensor is now underway in Europe for monitoring diabetics, according to Zhao. Volunteers eat sugary food to spike their glucose levels and prompt the monitor to register the changes.
"If this pans out, with approval from FDA, then consumers could get the sensors implanted in their core to measure their levels of glucose, oxygen, and lactate," Zhao said.
Lactate, especially, interests DARPA because it's a first responder molecule to the onset of trauma, sepsis, and potentially infection.
"The sensor could potentially detect rise of these [body chemistry numbers] and alert the user to prevent onset of dangerous illness."
2) NEAR INSTANTANEOUS VACCINE PROTECTION DURING A PANDEMIC
Traditional vaccines can take months or years to develop, then weeks to become effective once you get it. But when an unknown virus emerges, there's no time to waste.
This program, called P3, envisions a much more ambitious approach to stop a pandemic in its tracks.
"We want to confer near instantaneous protection by doing it a different way – enlist the body as a bioreactor to produce therapeutics," said Col. Matthew Hepburn, the program manager.
So how would it work?
To fight a pandemic, we will need 20,000 doses of a vaccine in 60 days.
If you have antibodies against a certain infection, you'll be protected against that infection. This idea is to discover the genetic code for the antibody to a specific pathogen, manufacture those pieces of DNA and RNA, and then inject the code into a person's arm so the muscle cells will begin producing the required antibodies.
"The amazing thing is that it actually works, at least in animal models," said Hepburn. "The mouse muscles made enough protective antibodies so that the mice were protected."
The next step is to test the approach in humans, which the program will do over the next two years.
But the hard part is actually not discovering the genetic code for highly potent antibodies, according to Hepburn. In fact, researchers already have been able to do so in two to four weeks' time.
"The hard part is once I have an antibody, a large pharma company will say in 2 years, I can make 100-200 doses. Give us 4 years to get to 20,000 doses. That's not good enough," Hepburn said.
To fight a pandemic, we will need 20,000 doses of a vaccine in 60 days.
"We have to fundamentally change the idea that it takes a billion dollars and ten years to make a drug," he concluded. "We're going to do something radically different."
3) RAPID DIAGNOSING OF PATHOGEN EXPOSURE THROUGH EPIGENETICS
Imagine that you come down with a mysterious illness. It could be caused by a virus, bacteria, or in the most extreme catastrophe, a biological agent from a weapon of mass destruction.
What if a portable device existed that could identify--within 30 minutes—which pathogen you have been exposed to and when? It would be pretty remarkable for soldiers in the field, but also for civilians seeking medical treatment.
This is the lofty ambition of a DARPA program called Epigenetic Characterization and Observation, or ECHO.
Its success depends on a biological phenomenon known as the epigenome. While your DNA is relatively immutable, your environment can modify how your DNA is expressed, leaving marks of exposure that register within seconds to minutes; these marks can persist for decades. It's thanks to the epigenome that identical twins – who share identical DNA – can differ in health, temperament, and appearance.

These three mice are genetically identical. Epigenetic differences, however, result in vastly different observed characteristics.
Reading your epigenetic marks could theoretically reveal a time-stamped history of your body's environmental exposures.
Researchers in the ECHO program plan to create a database of signatures for exposure events, so that their envisioned device will be able to quickly scan someone's epigenome and refer to the database to sort out a diagnosis.
"One difficult part is to put a timestamp on this result, in addition to the sign of which exposure it was -- to tell us when this exposure happened," says Thomas Thomou, a contract scientist who is providing technical assistance to the ECHO program manager.
Other questions that remain up in the air for now: Do all humans have the same epigenetic response to the same exposure events? Is it possible to distinguish viral from bacterial exposures? Does dose and duration of exposure affect the signature of epigenome modification?
The program will kick off in January 2019 and is planned to last four years, as long as certain milestones of development are reached along the way. The desired prototype would be a simple device that any untrained person could operate by taking a swab or a fingerprick.
"In an outbreak," says Dr. Thomou, "it will help everyone on the ground immediately to have a rapidly deployable machine that will give you very quick answers to issues that could have far-reaching ramifications for public health safety."
Kira Peikoff was the editor-in-chief of Leaps.org from 2017 to 2021. As a journalist, her work has appeared in The New York Times, Newsweek, Nautilus, Popular Mechanics, The New York Academy of Sciences, and other outlets. She is also the author of four suspense novels that explore controversial issues arising from scientific innovation: Living Proof, No Time to Die, Die Again Tomorrow, and Mother Knows Best. Peikoff holds a B.A. in Journalism from New York University and an M.S. in Bioethics from Columbia University. She lives in New Jersey with her husband and two young sons. Follow her on Twitter @KiraPeikoff.
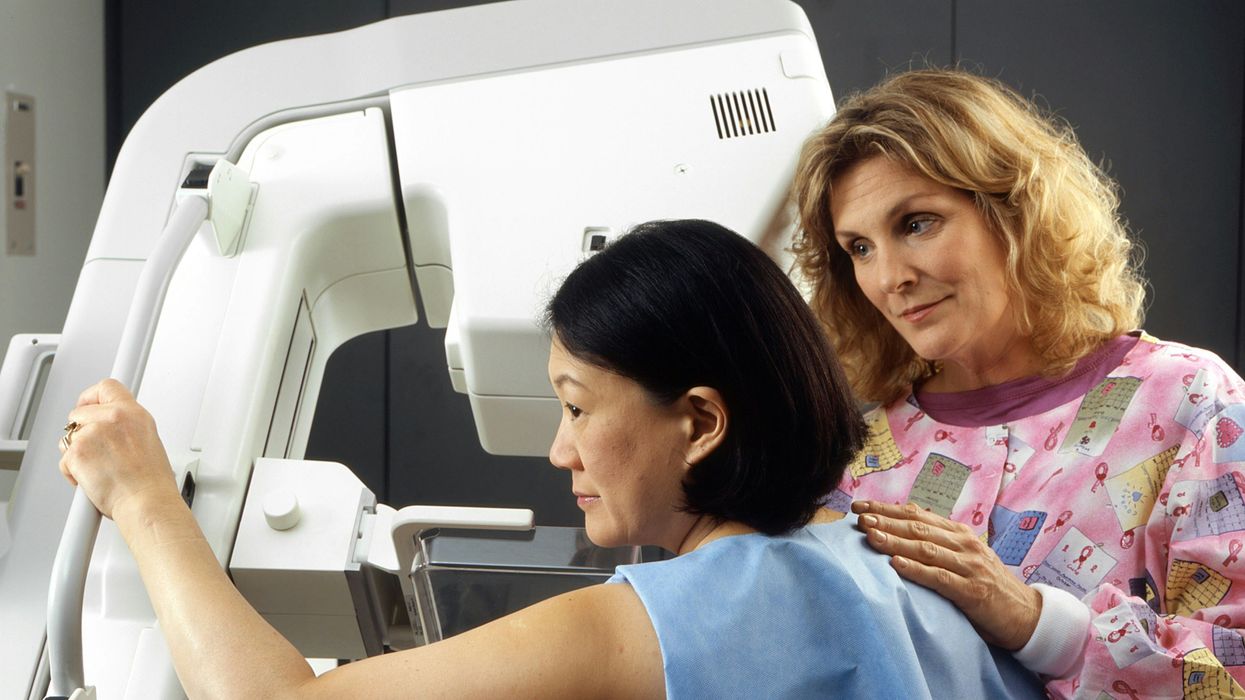
A woman receives a mammogram, which can detect the presence of tumors in a patient's breast.
When a patient is diagnosed with early-stage breast cancer, having surgery to remove the tumor is considered the standard of care. But what happens when a patient can’t have surgery?
Whether it’s due to high blood pressure, advanced age, heart issues, or other reasons, some breast cancer patients don’t qualify for a lumpectomy—one of the most common treatment options for early-stage breast cancer. A lumpectomy surgically removes the tumor while keeping the patient’s breast intact, while a mastectomy removes the entire breast and nearby lymph nodes.
Fortunately, a new technique called cryoablation is now available for breast cancer patients who either aren’t candidates for surgery or don’t feel comfortable undergoing a surgical procedure. With cryoablation, doctors use an ultrasound or CT scan to locate any tumors inside the patient’s breast. They then insert small, needle-like probes into the patient's breast which create an “ice ball” that surrounds the tumor and kills the cancer cells.
Cryoablation has been used for decades to treat cancers of the kidneys and liver—but only in the past few years have doctors been able to use the procedure to treat breast cancer patients. And while clinical trials have shown that cryoablation works for tumors smaller than 1.5 centimeters, a recent clinical trial at Memorial Sloan Kettering Cancer Center in New York has shown that it can work for larger tumors, too.
In this study, doctors performed cryoablation on patients whose tumors were, on average, 2.5 centimeters. The cryoablation procedure lasted for about 30 minutes, and patients were able to go home on the same day following treatment. Doctors then followed up with the patients after 16 months. In the follow-up, doctors found the recurrence rate for tumors after using cryoablation was only 10 percent.
For patients who don’t qualify for surgery, radiation and hormonal therapy is typically used to treat tumors. However, said Yolanda Brice, M.D., an interventional radiologist at Memorial Sloan Kettering Cancer Center, “when treated with only radiation and hormonal therapy, the tumors will eventually return.” Cryotherapy, Brice said, could be a more effective way to treat cancer for patients who can’t have surgery.
“The fact that we only saw a 10 percent recurrence rate in our study is incredibly promising,” she said.
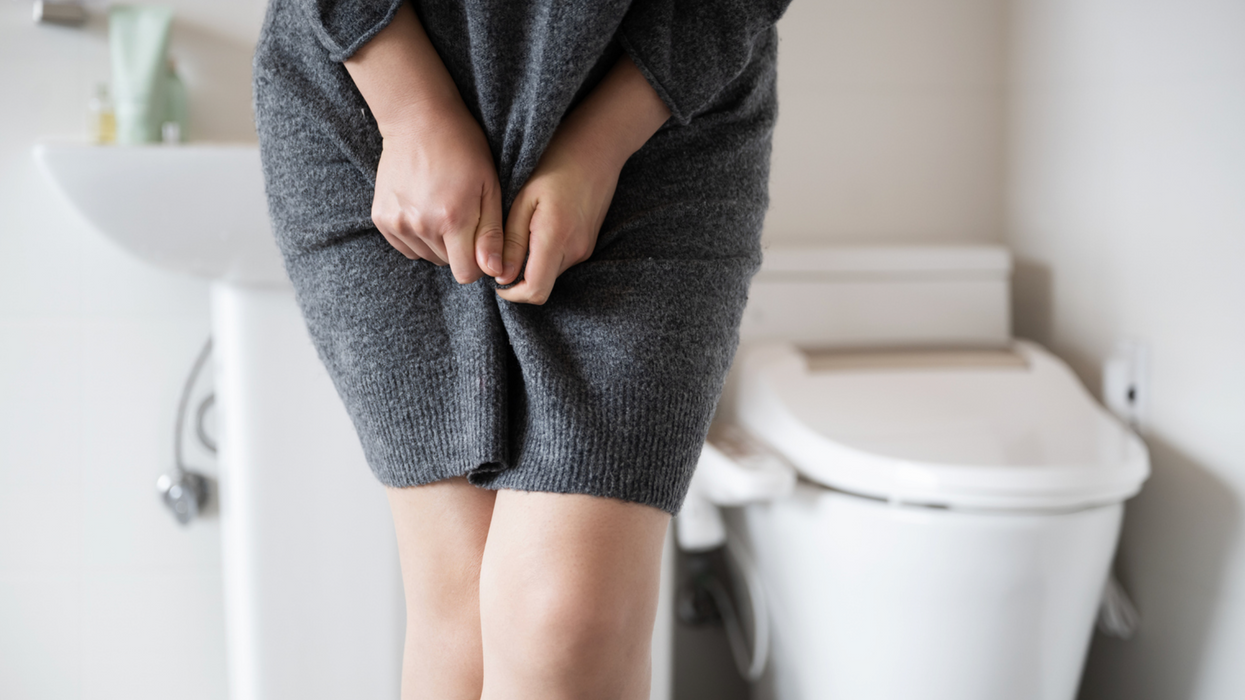
Urinary tract infections account for more than 8 million trips to the doctor each year.
Few things are more painful than a urinary tract infection (UTI). Common in men and women, these infections account for more than 8 million trips to the doctor each year and can cause an array of uncomfortable symptoms, from a burning feeling during urination to fever, vomiting, and chills. For an unlucky few, UTIs can be chronic—meaning that, despite treatment, they just keep coming back.
But new research, presented at the European Association of Urology (EAU) Congress in Paris this week, brings some hope to people who suffer from UTIs.
Clinicians from the Royal Berkshire Hospital presented the results of a long-term, nine-year clinical trial where 89 men and women who suffered from recurrent UTIs were given an oral vaccine called MV140, designed to prevent the infections. Every day for three months, the participants were given two sprays of the vaccine (flavored to taste like pineapple) and then followed over the course of nine years. Clinicians analyzed medical records and asked the study participants about symptoms to check whether any experienced UTIs or had any adverse reactions from taking the vaccine.
The results showed that across nine years, 48 of the participants (about 54%) remained completely infection-free. On average, the study participants remained infection free for 54.7 months—four and a half years.
“While we need to be pragmatic, this vaccine is a potential breakthrough in preventing UTIs and could offer a safe and effective alternative to conventional treatments,” said Gernot Bonita, Professor of Urology at the Alta Bro Medical Centre for Urology in Switzerland, who is also the EAU Chairman of Guidelines on Urological Infections.
The news comes as a relief not only for people who suffer chronic UTIs, but also to doctors who have seen an uptick in antibiotic-resistant UTIs in the past several years. Because UTIs usually require antibiotics, patients run the risk of developing a resistance to the antibiotics, making infections more difficult to treat. A preventative vaccine could mean less infections, less antibiotics, and less drug resistance overall.
“Many of our participants told us that having the vaccine restored their quality of life,” said Dr. Bob Yang, Consultant Urologist at the Royal Berkshire NHS Foundation Trust, who helped lead the research. “While we’re yet to look at the effect of this vaccine in different patient groups, this follow-up data suggests it could be a game-changer for UTI prevention if it’s offered widely, reducing the need for antibiotic treatments.”